Stat 437 Lecture Notes 2b
Xiongzhi Chen
Washington State University
Visualization via ggplot2: adjusting scales
Overview
We will cover how to “(c) manually set some scales” for a plot, focusing on
- Position scales
- Colour scales
- Manual scales
The contents for (c) are based on Chapters 5 and 6 of the book “ggplot2: elegant graphs for data analysis” by Hadley Wickham.
Overview
Scales
- control mapping from data to aesthetics (e.g., size, colour, position, or shape)
- provide
guides (e.g., axes and legends) - can be roughly divided into four categories: position scales, colour scales, the manual discrete scale, and the identity scale; for position aesthetics, axes are guides; for all other aesthetics, legends do the job
A scale is needed for each plot, and ggplot2 will add a default scale when none is specified by a user.
Four categories of scales
- “Position scales” are used to map continuous and discrete variables onto the plotting region and construct the corresponding axes
- “Colour scales” are used to map continuous and discrete variables to colours
- “Manual scales” are used to map discrete variables to a user’s choice of symbol size, line type, shape, or colour, and create the corresponding legend
- “The identity scale” is used to plot variable values directly to the aesthetic rather than mapping them (to some other aesthetics)
Position scales
Set ranges for an axis
# set ranges for both x-axis and y-axis
lims(...)
# set range for x-axis
xlim(...)
# set range for y-axis
ylim(...)A remark
By default, the limits of position scales extend (or expand) a little past the range of data. This ensures that data do not overlap axes.
One can control the amount of expansion with the expand argument. This parameter should be a numeric vector of length two. The first element gives the multiplicative expansion, and the second the additive expansion.
If no expansion is needed, use
scale_x_continuous(expand=c(0,0))Base plot
Plot cty (mpg in city) vs hwy (mpg on highway):
> p =ggplot(mpg)+geom_point(aes(x=cty,y=hwy,color=class)); p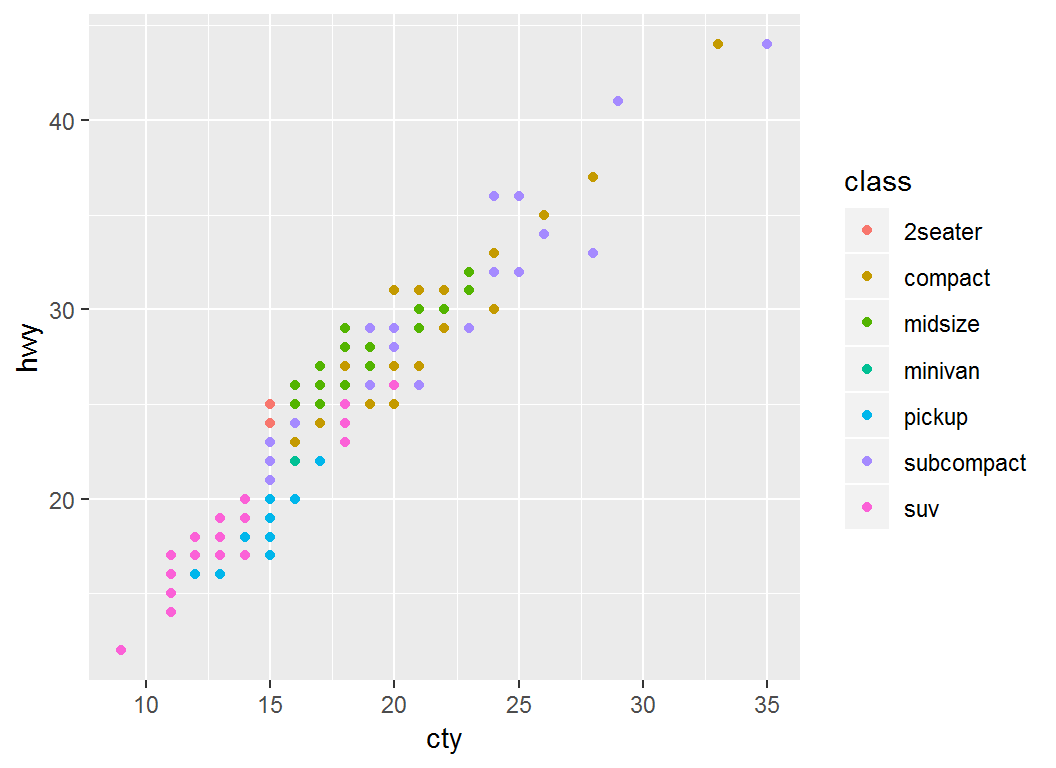
Set range for x-axis
> p + xlim(c(4,20))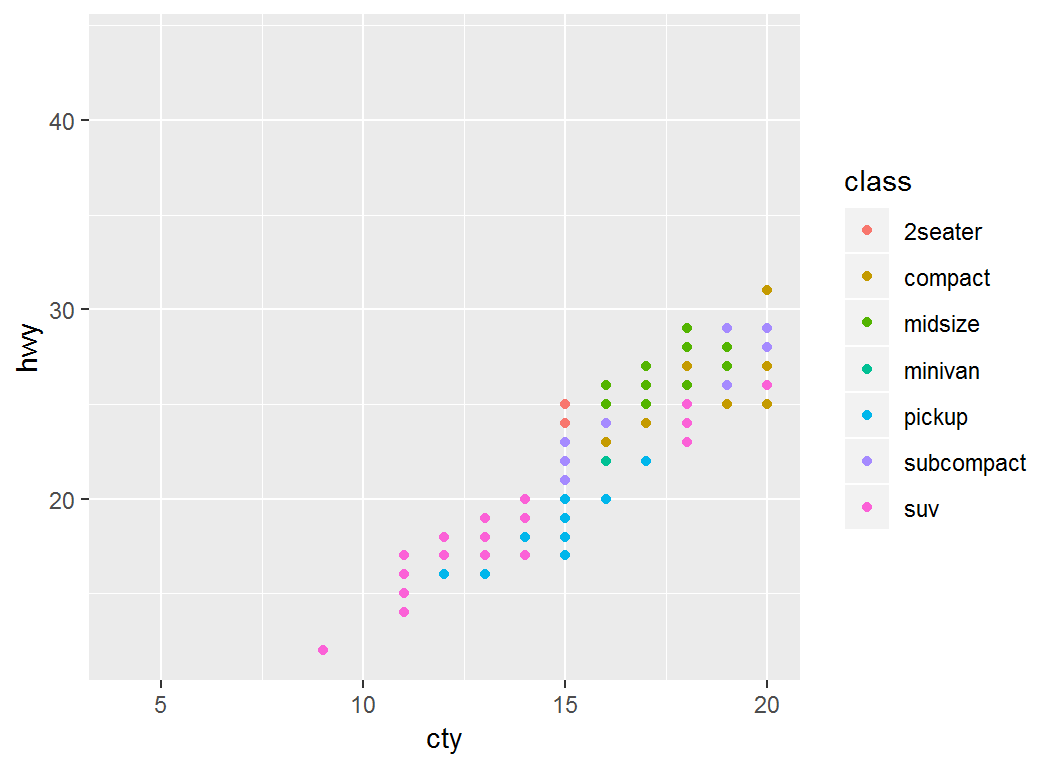
> # use xlim(c(NA,20)) to set an automatic lower limitSet range for both axes
> p + lims(x = c(10, 20), y = c(3, 5))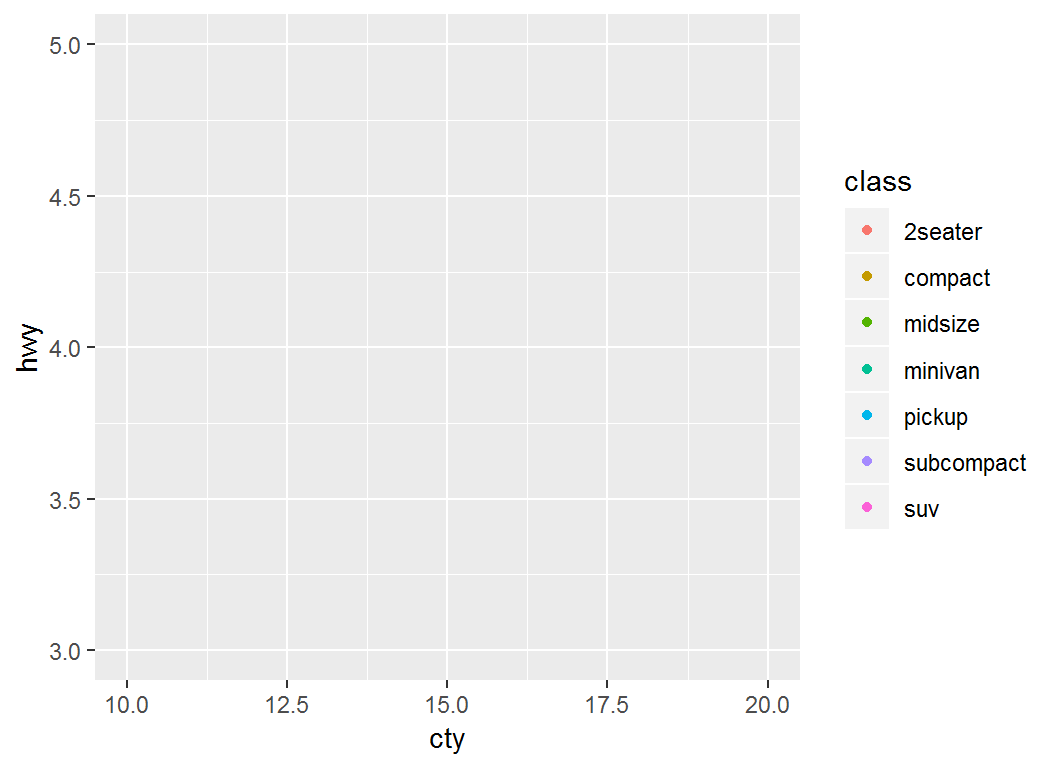
scale_*_continuous
scale_x_continuous (or scale_y_continuous) controls the x (or y) axis for continuous variables, and often sets breaks, labels, na.value, and/or trans:
breaks: a numeric vector of tick positionslabels: a character vector giving labels (must be same length asbreaks)na.value=value: missing values are set asvaluetrans: transformations such asscale_*_log10(),scale_*_sqrt()andscale_*_reverse()
Illustration: base plot
> # Plot `displ` vs `hwy`:
> p1 = ggplot(mpg, aes(displ,hwy)) + geom_point(); p1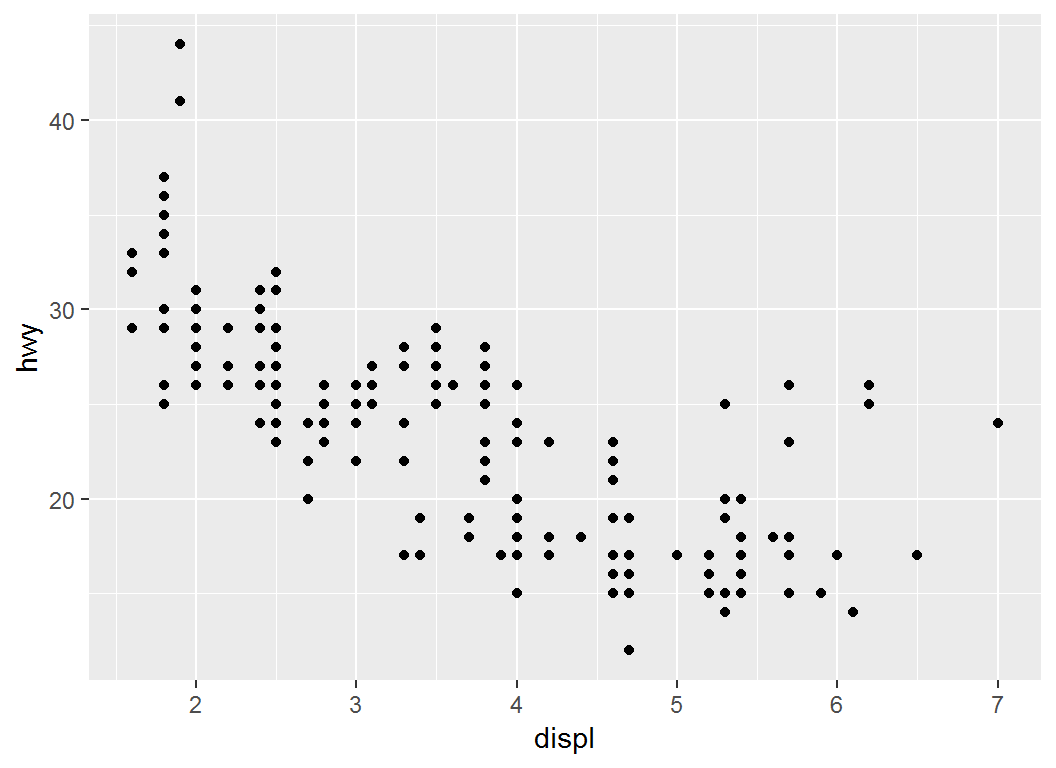
Illustration: breaks
> # choose where the x-axis ticks appear
> p1 + scale_x_continuous(breaks = c(2, 4, 6))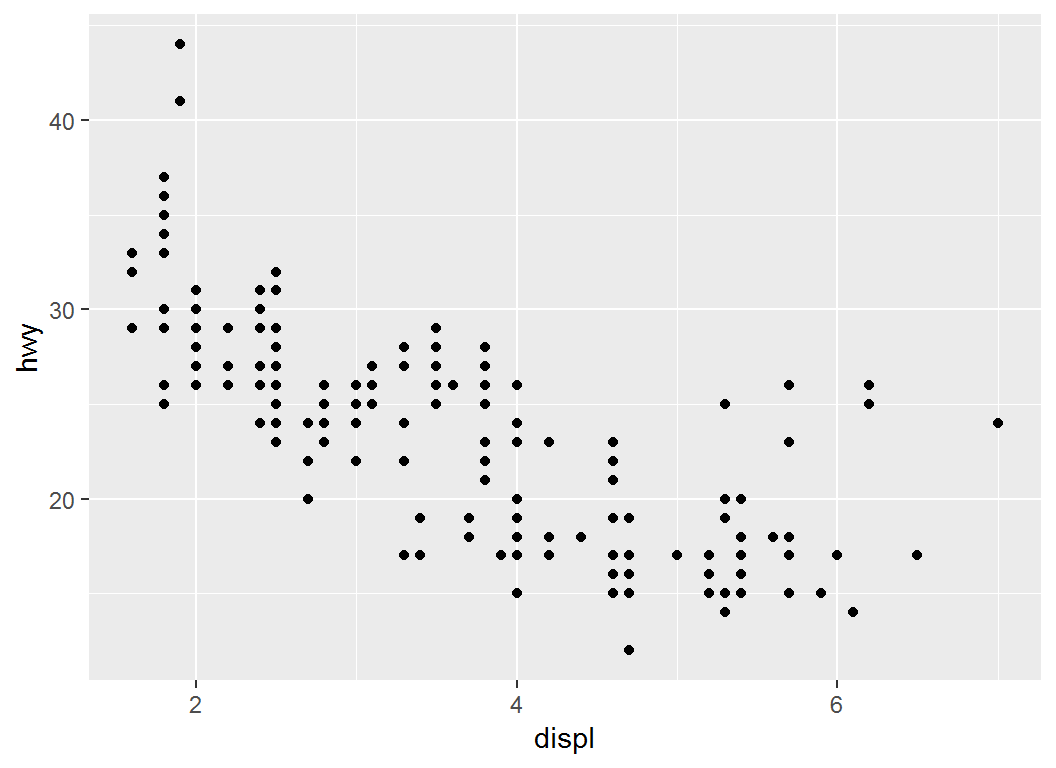
Illustration: label
> # personalized labels for ticks at specified positions
> p1 + scale_x_continuous(breaks = c(2, 4, 6),
+ label = c("two", "four", "six"))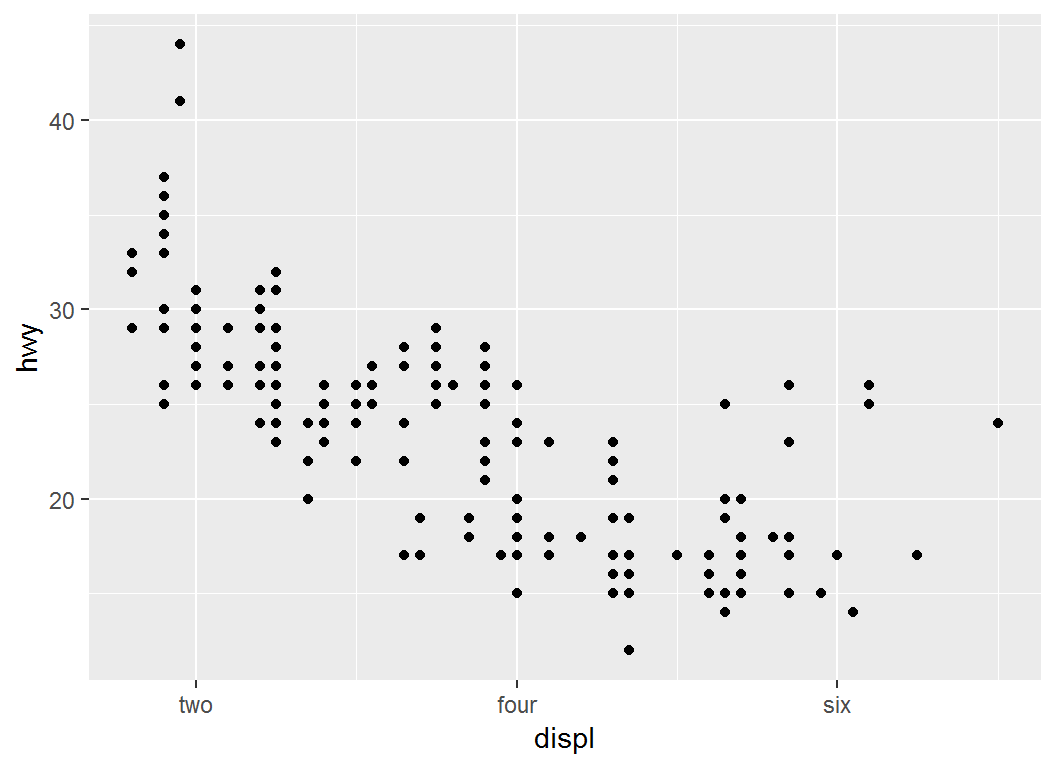
Illustration: trans
> # y-axis on natural logarithmic scale via `trans=log`
> p1 + scale_y_continuous(trans = "log")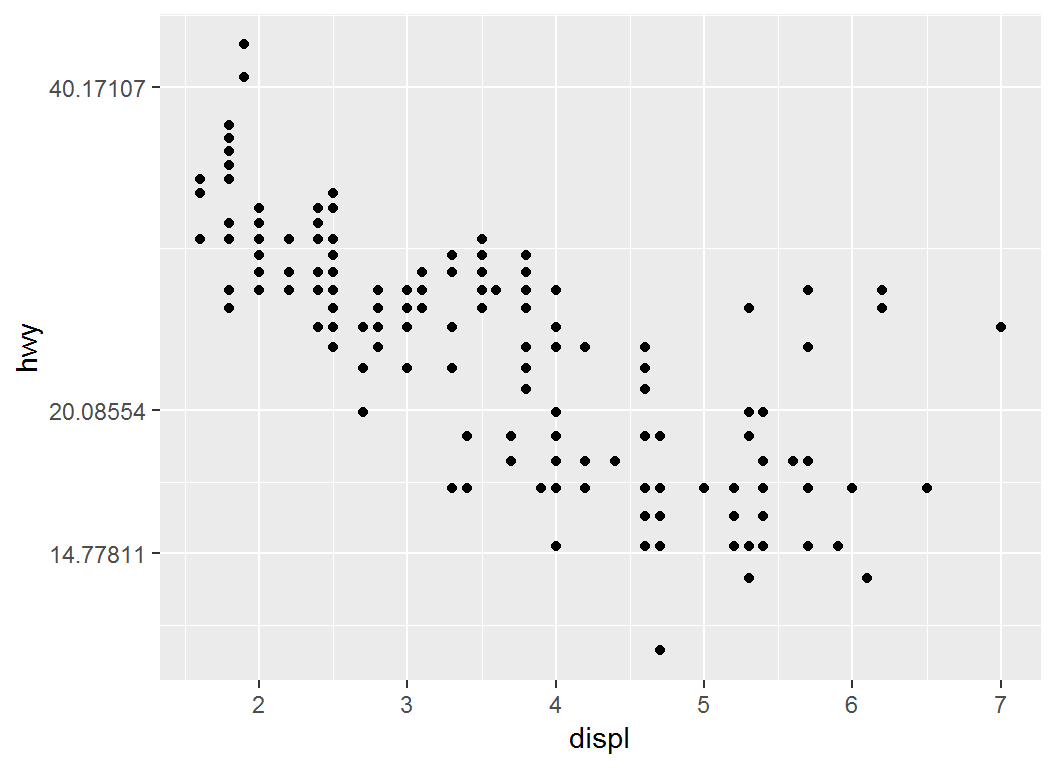
trans: options
Table 6.2 from book “ggplot2”
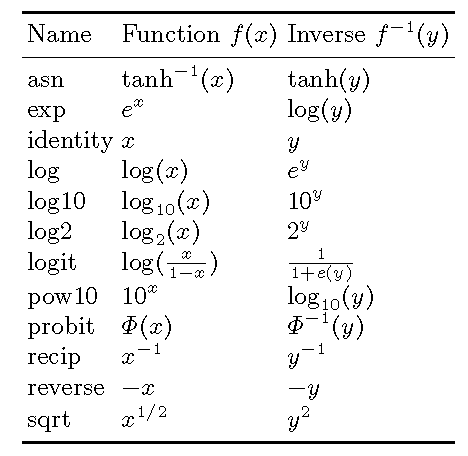

scale_*_discrete
scale_x_discrete (or scale_y_discrete):
- controls the
x(ory) axis for discrete variables - is often used to set
breaks,labels,na.value, and/ortrans. - has syntax and usage similar to those of
scale_x_continuous(andscale_y_continuous)
Illustration
Base layer: bar plot for drv:
> p = ggplot(mpg, aes(x = drv)) + geom_bar(); p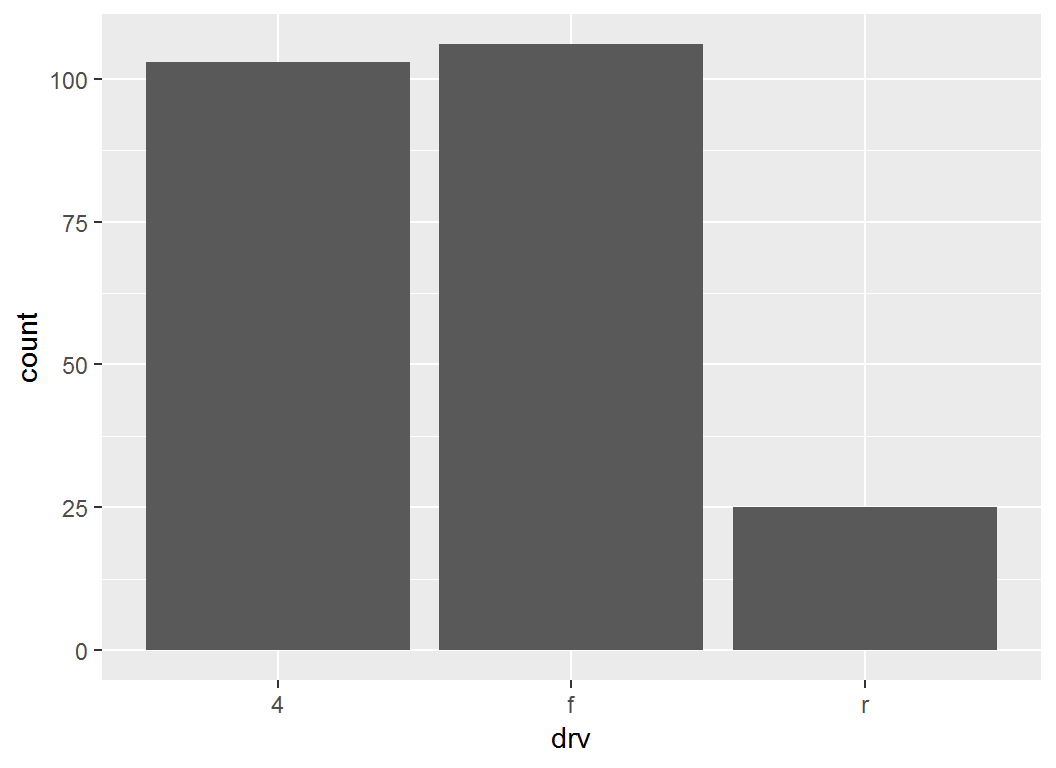
Illustration
> # re-label x-axis ticks
> p + scale_x_discrete(labels =
+ c("4 wheel drive", "front drive", "rear drive"))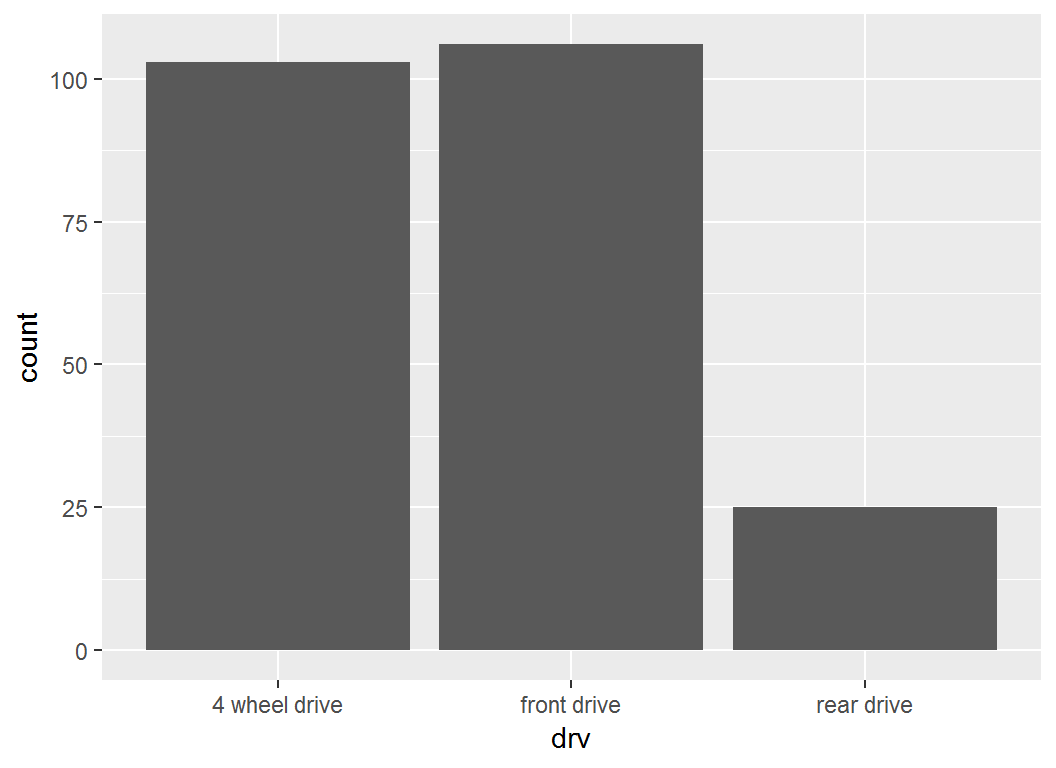
Colour scales for continuous variables
Overview
After position, probably the most commonly used aesthetic is colour. For this aesthetic and continuous variables, there are three methods, based on their gradient schemes:
scale_*_gradient()scale_*_gradient2()scale_*_gradientn())
Note: colour is exchangeable with color
Method I: Two-colour gradient
scale_colour_gradient() and scale_fill_gradient():
- each being a two-colour gradient “low-high”, i.e., with a low end and a high end
- arguments
low(for “low end” ) andhigh(for “high end”) control the colours at the low end and high end of the gradient, respectively
Method II: Diverging-colour gradient
scale_colour_gradient2() and scale_fill_gradient2():
- each being a three-colour gradient “low-mid-high”, i.e., with a low, mid, and high end
- each having a
midcolour for the colour ofmidpoint midpointdefaults to \(0\) but can be set to any value
These two functions are particularly useful for creating diverging colour schemes
Method III: n-colour gradient
scale_colour_gradientn() and scale_fill_gradientn():
- each being a custom
n-colour gradient - each requiring a vector of colours in the
coloursargument; by default, these colours will be evenly spaced along the range of the data
Illustration
> # Plot `cty` vs `hwy` with default scheme `color=displ`
> p = ggplot(mpg)+geom_point(aes(cty,hwy,color=displ))
> p # note legend title "displ""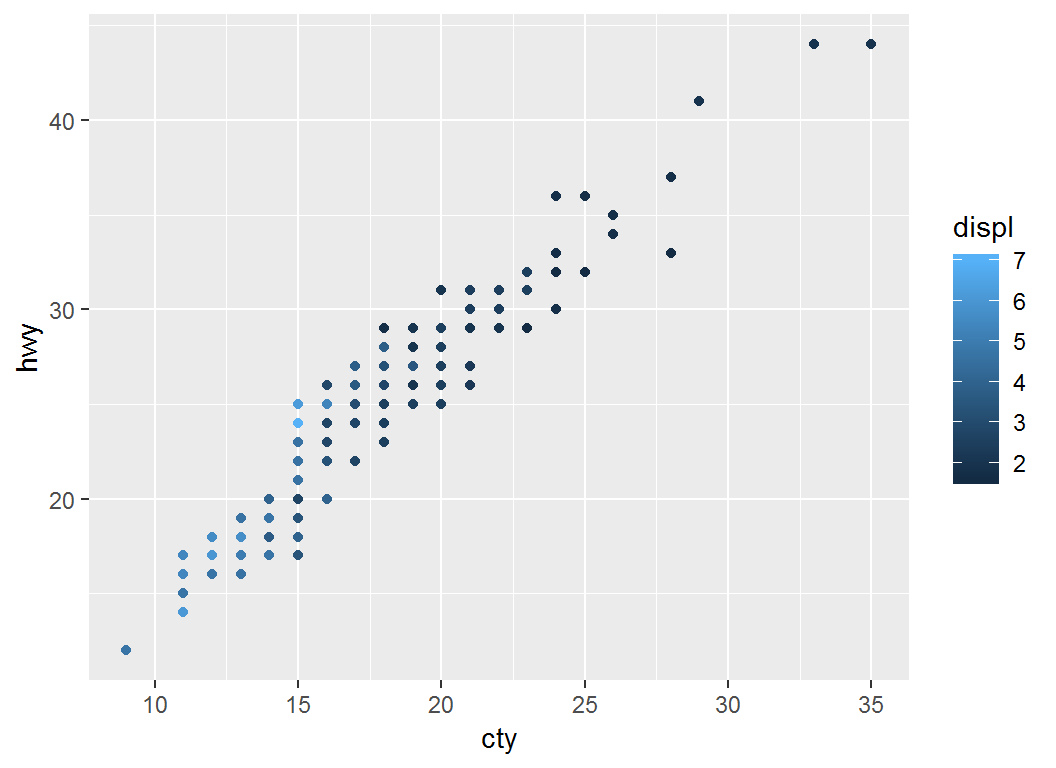
Adjust coloring: low-mid-high
> p2a1=p + scale_colour_gradient2("Displacement",low="gray",
+ mid="blue",high="red",midpoint=mean(mpg$displ))
> p2a1 # note legend title "Displacement" and `midpoint`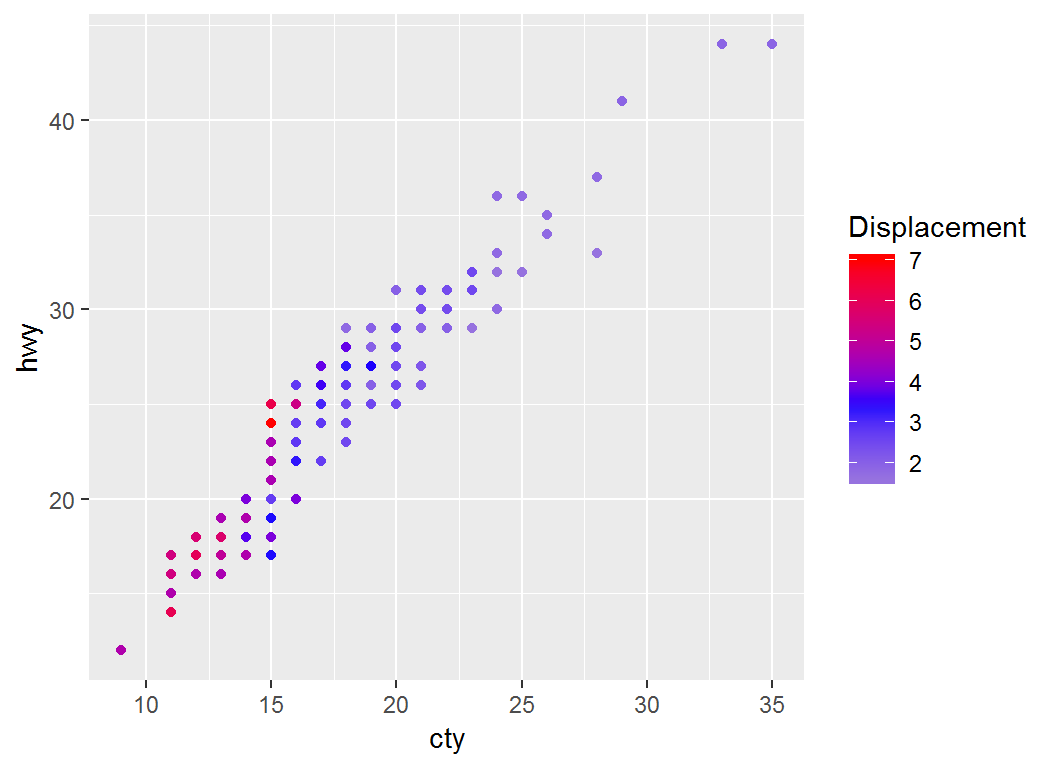
Remark on midpoint
> p2a2 = p+scale_colour_gradient2("Displacement",low="gray",
+ mid="blue",high="red",midpoint=2*mean(mpg$displ))
> library(gridExtra); grid.arrange(p2a1,p2a2,nrow=2)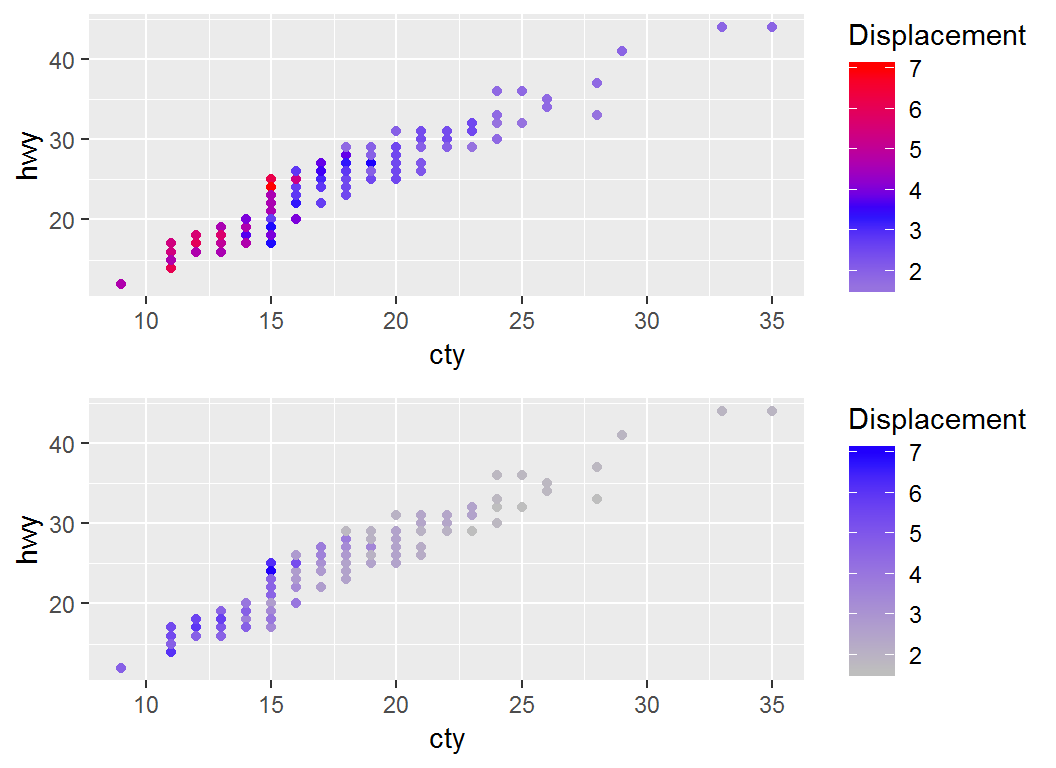
Visualise 3D surfaces in 2D
The faithful dataset (in library MASS) records waiting times (waiting) between eruptions and eruption times in minutes eruption for the Old Faithful geyser in Yellowstone Park:
> library(MASS); head(faithful)
eruptions waiting
1 3.600 79
2 1.800 54
3 3.333 74
4 2.283 62
5 4.533 85
6 2.883 55Density function \(z=f(x,y)\) for \((x,y)\)=(eruptions,waiting) can be visualized via 2D contours
Obtain estimated density
Obtain density estimate for (eruptions,waiting):
- load library
MASS; applykde2d, i.e., 2D kernel density estimation, tofaithfuldata set - data frame “f2d” has 3 columns x, y and z, where z is the value of the estimated density evaluated at (x,y)
> f2d <- with(faithful, MASS::kde2d(eruptions, waiting,
+ h = c(1, 10), n = 50))
> df <- with(f2d, cbind(expand.grid(x, y), as.vector(z)))
> names(df) <- c("eruptions", "waiting", "density")
> head(df)
eruptions waiting density
1 1.600000 43 0.003216159
2 1.671429 43 0.004146406
3 1.742857 43 0.004987802
4 1.814286 43 0.005611508
5 1.885714 43 0.005921813
6 1.957143 43 0.005882327Visualize density estimate
> erupt <- ggplot(df,aes(waiting,eruptions,fill = density))+
+ geom_tile()+
+ scale_x_continuous(expand = c(0, 0))+
+ scale_y_continuous(expand = c(0, 0))Note the use of
geom_tile()andfill = densityscale_*_continuous(expand = c(0, 0))
Visualize density estimate
> erupt #max(df$density)=0.037, min(df$density)=10^(-24)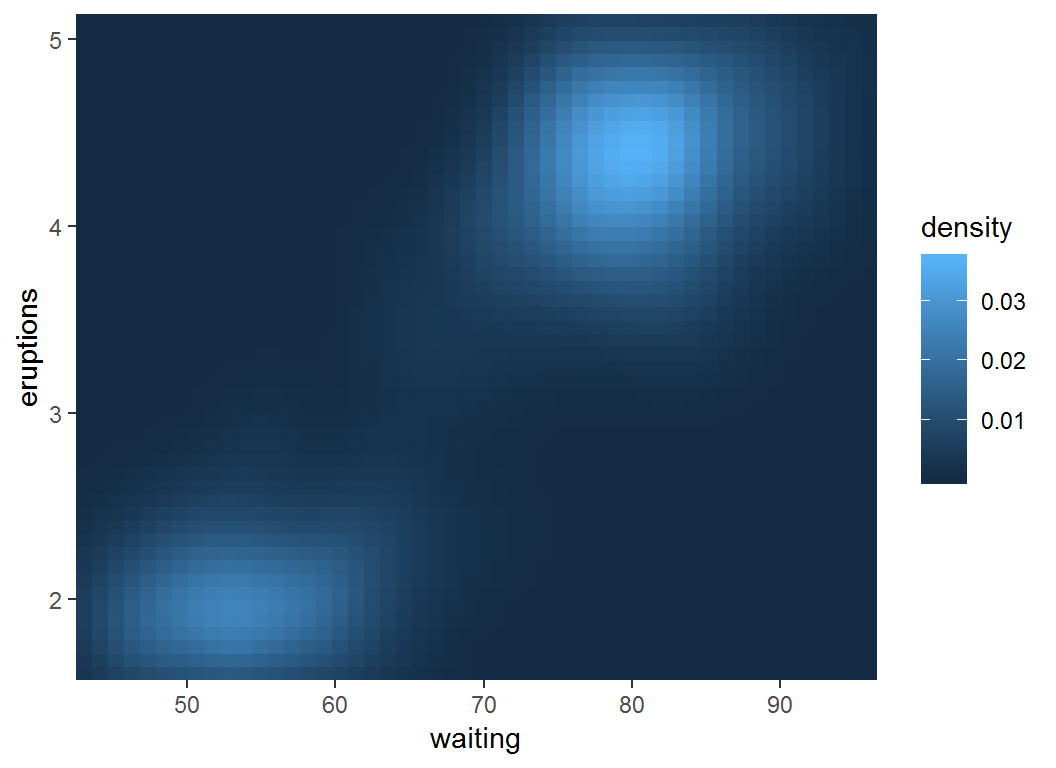
Adjust coloring: low-high
> # `limits = c(0, 0.04)` sets range for values in legend
> erupt + scale_fill_gradient(limits = c(0,0.04),
+ low = "white", high = "black")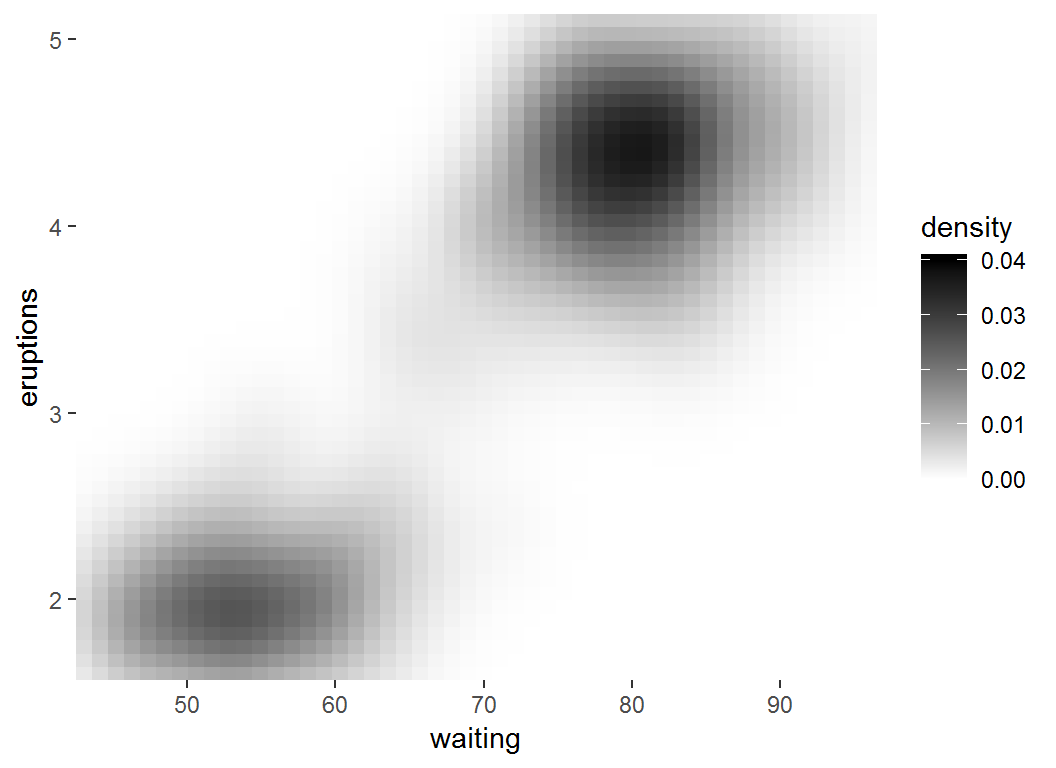
Adjust coloring: low-mid-high
> erupt + scale_fill_gradient2(limits = c(0, 0.04),
+ midpoint = mean(df$density))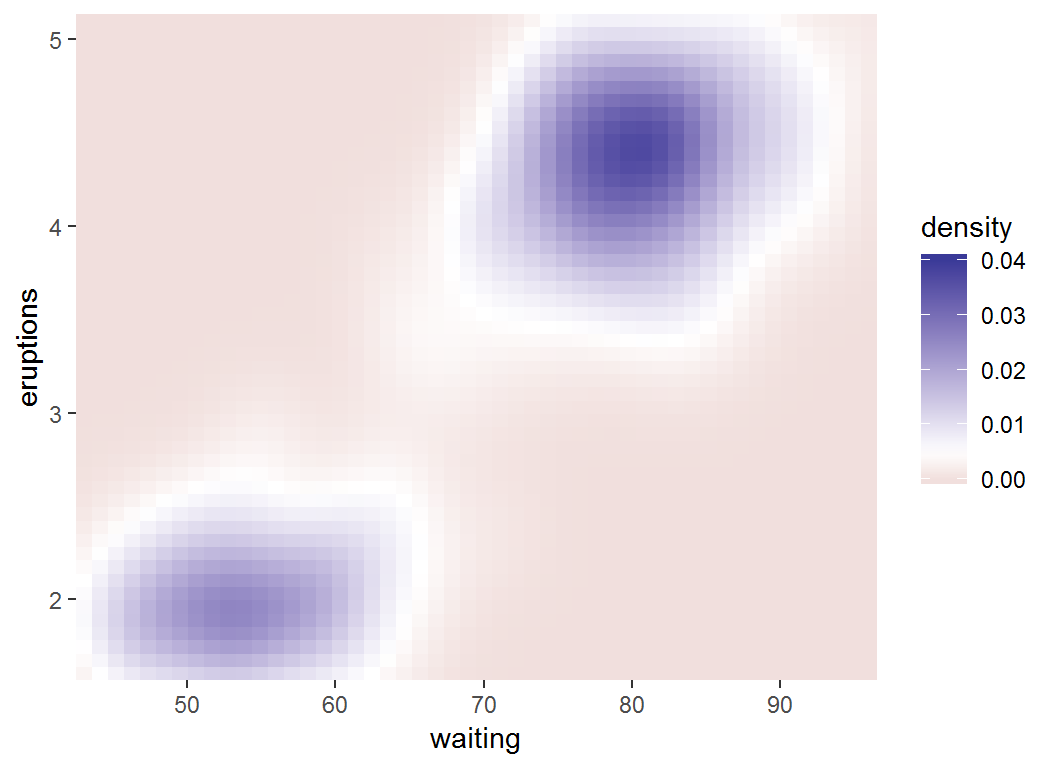
Remarks
Pay attention to how the range for
densityin the legend is controlled bylimitsinscale_fill_gradientorscale_fill_gradient2There are palettes available for color scales
Color scales for discrete variables
Overview
Two methods for colour scales for discrete data:
- choosing evenly spaced colors frm the color wheel (via, e.g.,
scale_colour_hue());scale_colour_hue()works well for up to about eight colours - selecting colors from hand-picked sets (via , e.g.,
RColorBrewer)
Popular palettes of RColorBrewer are “Set1” and “Dark2” for points and “Set2”, “Pastel1”, “Pastel2” and “Accent” for areas. RColorBrewer::display.brewer.all() lists all palettes.
Illustration
Part of msleep data set (from library ggplot2):
# A tibble: 6 x 3
brainwt bodywt vore
<dbl> <dbl> <chr>
1 NA 50 carni
2 0.0155 0.48 omni
3 NA 1.35 herbi
4 0.00029 0.019 omni
5 0.423 600 herbi
6 NA 3.85 herbibrainwt(brain weight in kilograms);bodywt(body weight in kilograms);vore(carnivore, omnivore or herbivore)
Illustration
Plot brainwt vs bodywt and color “point” by vore:
> p4 = ggplot(msleep)+
+ geom_point(aes(brainwt,bodywt,colour = vore))+
+ scale_x_continuous(trans="log")+
+ scale_y_continuous(trans="log")Note: both axes on nautral logarithmic scale
Illustration
> p4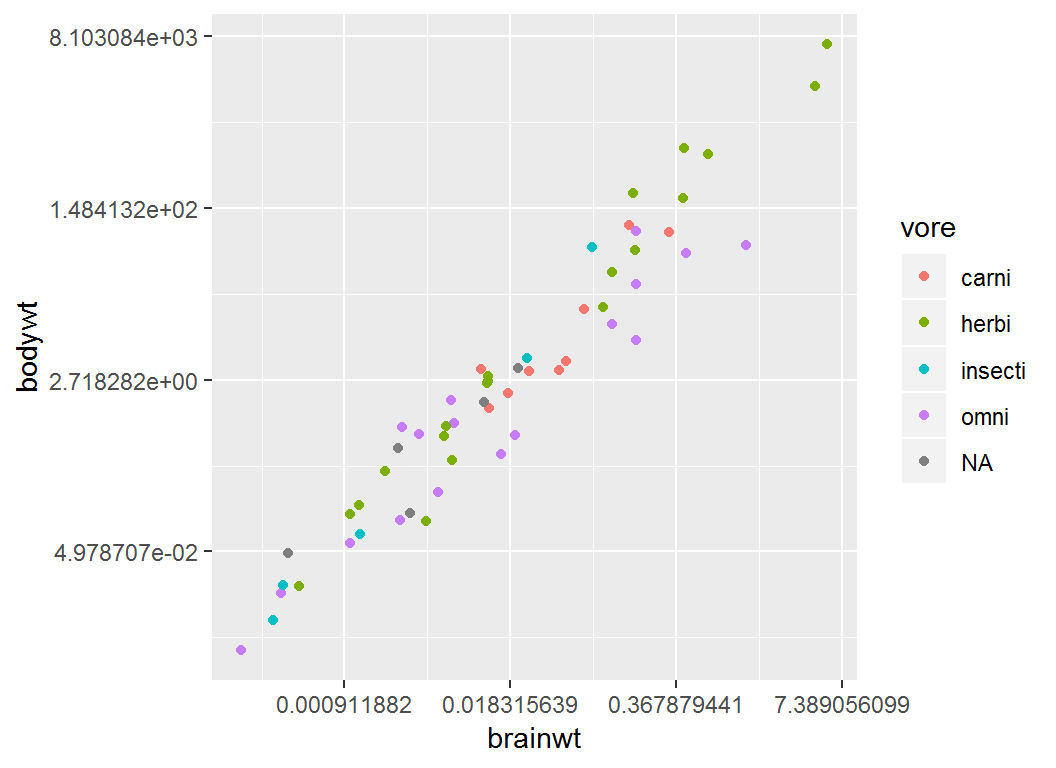
Adjust coloring by brewer
> p4 + scale_colour_brewer(palette = "Set1")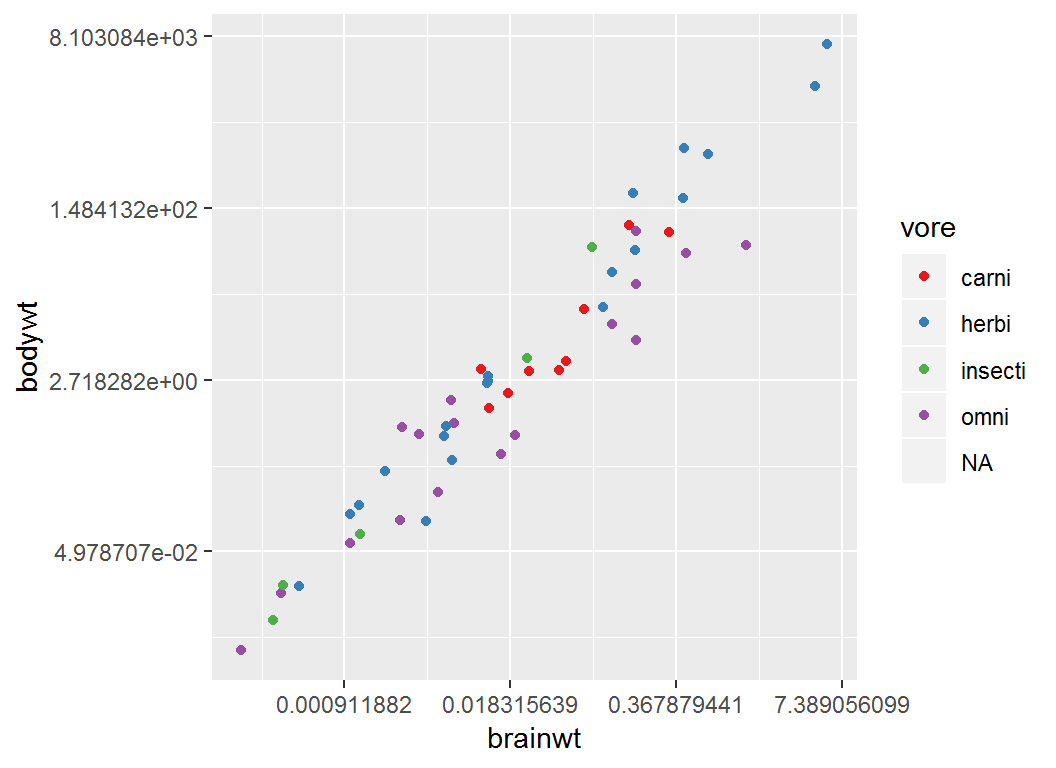
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 7 x64 (build 7601) Service Pack 1
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] MASS_7.3-49 knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] stringr_1.3.1 xfun_0.4 digest_0.6.18
[13] evaluate_0.12