Stat 437 Lecture Notes 3a
Xiongzhi Chen
Washington State University
Hierarchical clustering (HC): overview
Recap on K-means
To implement K-means clustering, a user needs to specify
- the number of clusters to obtain
- initial centers for clusters to start the optimization process
However, it is not easy to accurately estimate the true number of clusters in data
Overview on HC
Hierarchical clustering
- does not have the two disadvantages of K-means
- requires a measure of dissimilarity between (disjoint) groups (or clusters) of observations
- often produces hierarchical clusters, where each observation is a cluster at the finest level, clusters with increasing dissimilarities are nested, and all observations form one cluster at the coarsest level
However, users need to determine which level of hierarchy is sensible to obtain the eventual clusters
Two modes of HC
Two modes of hierarchical clustering
- “bottom-up” (or “agglomerative”): start with the finest level, merge two clusters with the smallest intergroup dissimilarity, and keep iterating until one cluster is obtained
- “top-down” (or “divisive”): start with the coarsest level, split the cluster into two groups with the largest intergroup dissimilarity, and keep iterating until each observation is a cluster
We will focus on “bottom-up”" (or “agglomerative”) HC
Illustration by simulated data
- 45 observations \(x_i,i=1,\ldots,45\)
- each observation \(x_i\) is 2-dimensional vector
- the 45 observations form 3 distinct classes
Note: distinct class labels will be indicated by distinct colors
Illustration by simulated data
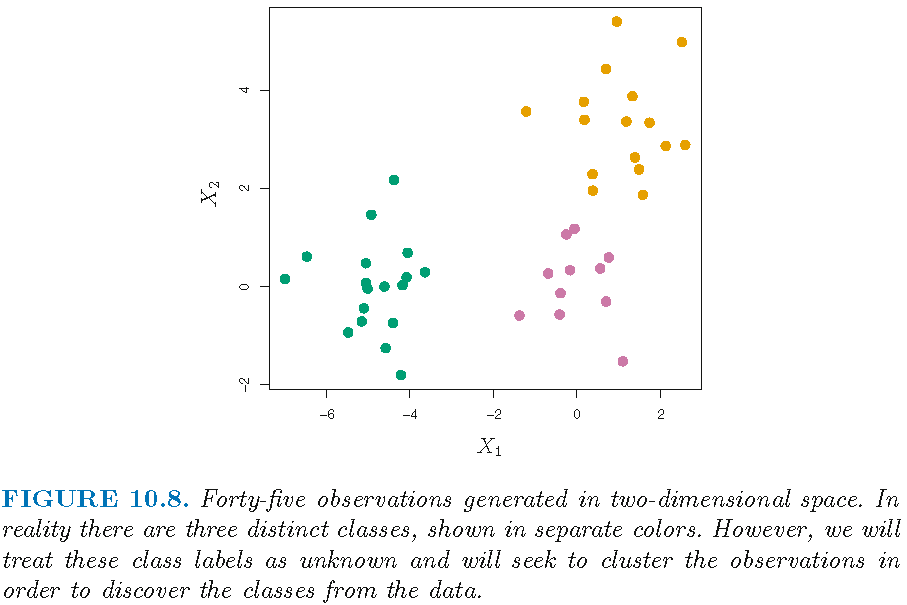
Illustration by simulated data
Suppose we do not know the class labels and perform hierarchical clustering by using
- Euclidean distance as the dissimilarity measure between each pair of observations
- complete linkage for disjoint groups that is based on Euclidean distance (to be discussed later)
We obtain a dendrogram that represents hierarchical clustering results as a tree-like structure
Illustration by simulated data
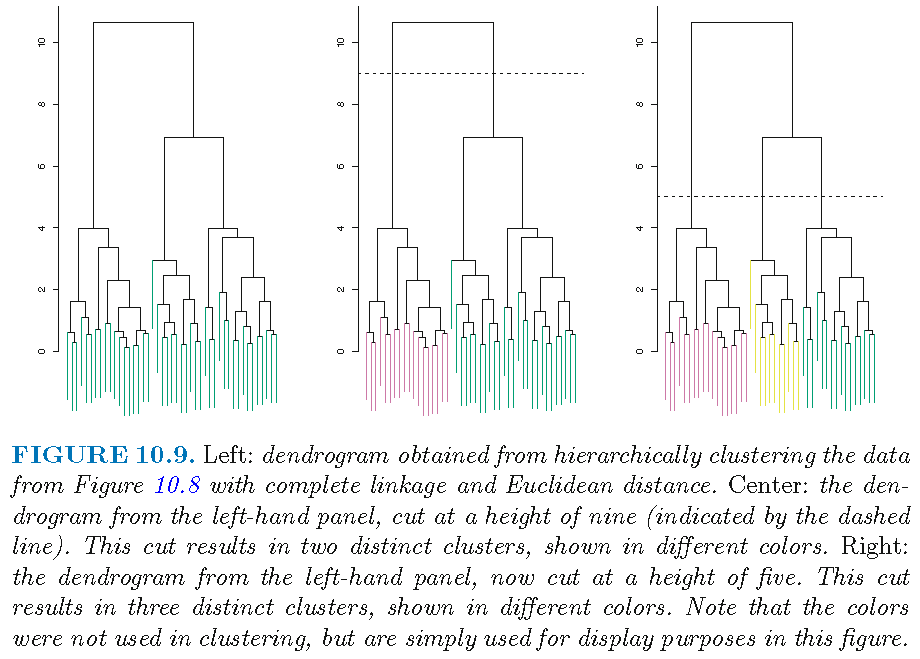
Understanding a dendrogram
A dendrogram is an upside-down tree with a height bar, where
- clusters at the bottom of the tree are leafs, where each leaf is an observation and a cluster
- as we move up the tree from the bottom, leafs begin to fuse into branches, and leafs and branches into branches
- the dissimilarity between two clusters indicates the height in the dendrogram at which these two clusters should be fused
- the larger the height is, the more dissimilar the branches or leafs are from each other at this height
Understanding a dendrogram
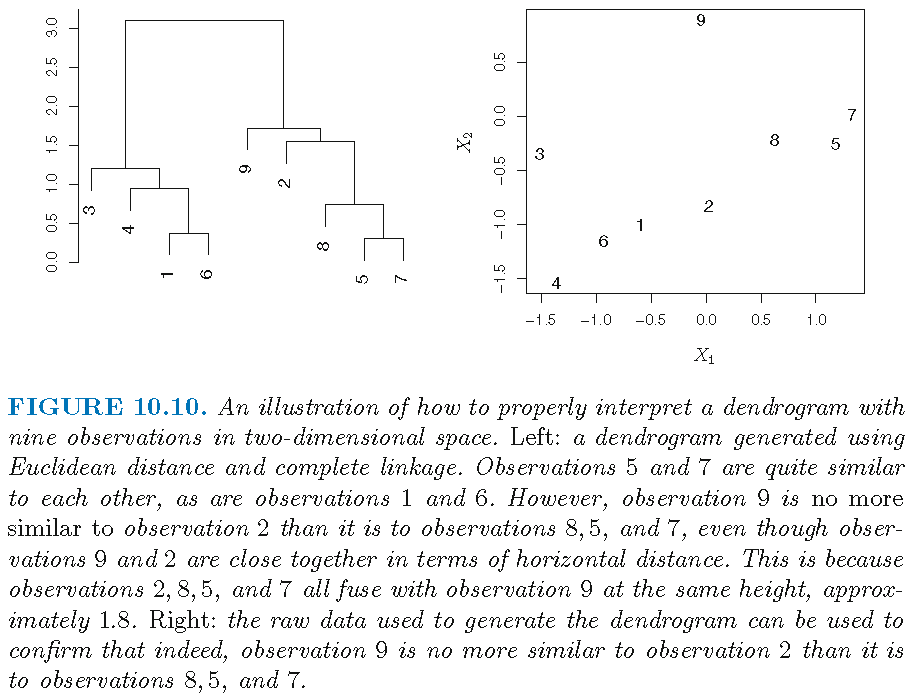
Understanding a dendrogram
When hierarchical clustering is represented by a dendrogram,
- the degree of similarity between two clusters is not determined by the horizontal distance between them on the dendrogram
- switching the left-right orientation of clusters at a given height that are subclusters of the same, immediate parent cluster does not change hierarchical clustering results
Caution on hierarchical clusters
Hierarchical clustering often produces hierarchical clusters, where clusters obtained by cutting the dendrogram at a given height (i.e., clusters at a given height) are necessarily nested within the clusters obtained by cutting the dendrogram at any greater height (i.e., clusters at any greater height), when there is only one criterion that determines clusters or groups in the data
However, hierarchical clustering will not necessarily be produced when sets of clusters are produced by different criteria
Caution on hierarchical clusters
Suppose we have observations from a group of people
- with a 50-50 split of males and females, giving the best two-group clustering that is obtained by gender
- evenly split among Americans, Japanese, and French, giving the best three-group clustering that is obtained by nationality
So, the two sets of groups are not nested, in the sense that the best 3-group clustering cannot be obtained splitting one group of the best 2-group clustering (since the group of people with the same nationality must contain people from both genders), and hence cannot be well represented by hierarchical clustering
Hierarchical clustering (HC): methodology and algorithm
Settings
- Assume there are \(p\) features \(x_j, j=1,\ldots,p\)
- Assume \(x_{ij},i=1,\ldots,n\) are \(n\) observations for the \(j\)th feature \(x_j\)
- Let \(\boldsymbol{x}_i = (x_{i1},\ldots,x_{ip}) \in \mathbb{R}^p\), i.e., \(\boldsymbol{x}_i\) is the \(i\)th observation for all \(p\) features
Using some dissimilarity measure, hierarchical clustering will split the \(n\) observations \(\boldsymbol{x}_i\) into sets of clusters at different dissimilarity levels (i.e., at different heights)
Agglomerative HC: algorithm
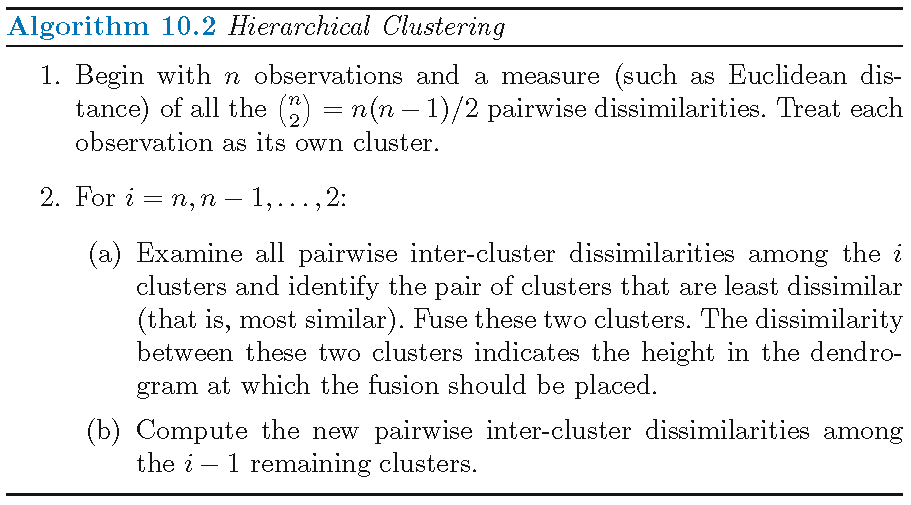

Dissimilarity meaures
The choice of a dissimilarity measure is crucial to hierarchical clustering, and there are many choices for this measure:
- Dissimilarity measure for each pair of observations (i.e., pairwise dissimilarity): Euclidean distance, the absolute value of sample correlation
- Dissimilarity measure for each pair of clusters (i.e., inter-cluster dissimilarity or linkage): complete linkage, single linkage, average linkage, centroid
Note: inter-cluster dissimilarity and pairwise dissimilarity should be compatible with each other in order to obtain hierarchical clustering
HC algorithm: illustration
Agglomerative HC with Euclidean distance and complete linkage:
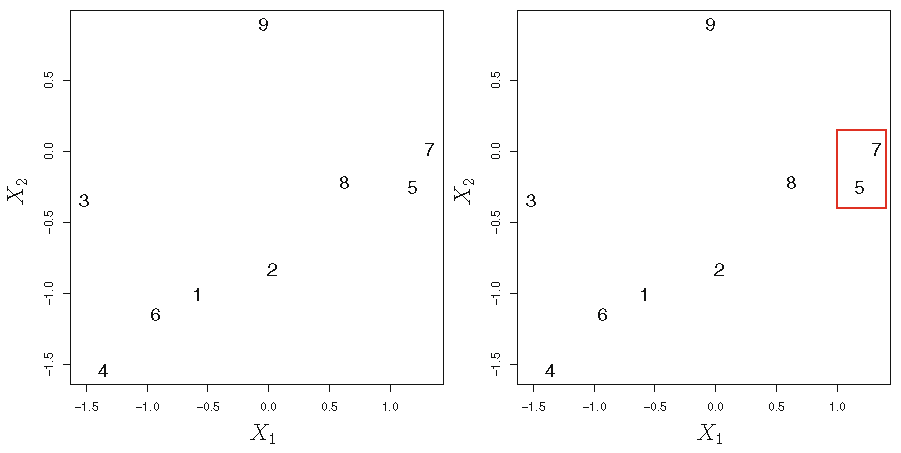
HC algorithm: illustration
Agglomerative HC with Euclidean distance and complete linkage:
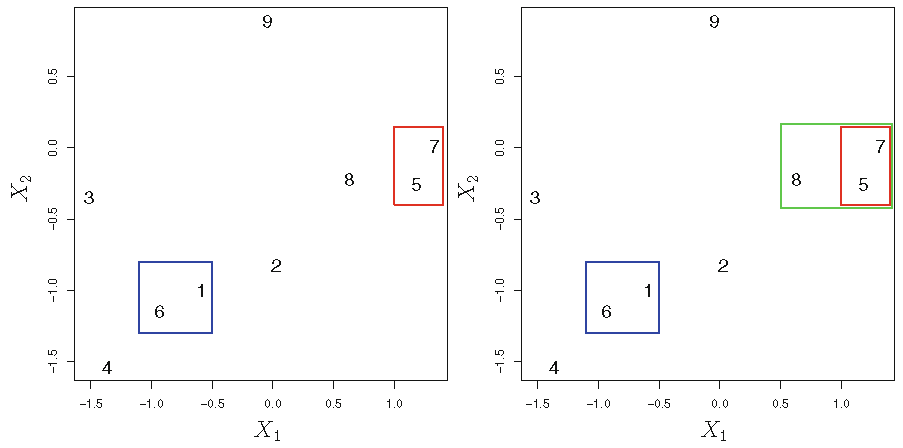
Different linkages
Single linkage, i.e., minimal intercluster dissimilarity: compute all pairwise dissimilarities between the observations in cluster A and the observations in cluster B, and record the smallest of these dissimilarities. Single linkage can result in extended, trailing clusters in which single observations are fused one-at-a-time
Centroid: dissimilarity between the centroid for cluster A and the centroid for cluster B. Centroid linkage may induce inversion in clustering, where two clusters are fused at a height below either of the individual clusters in the dendrogram
Different linkages
Average linkage, i.e., mean intercluster dissimilarity: compute all pairwise dissimilarities between the observations in cluster A and the observations in cluster B, and record the average of these dissimilarities
Complete linkage, i.e., maximal intercluster dissimilarity: compute all pairwise dissimilarities between the observations in cluster A and the observations in cluster B, and record the largest of these dissimilarities
Note: Average and complete linkages are generally preferred over single linkage, since they tend to yield more balanced dendrograms
Effects of linkage
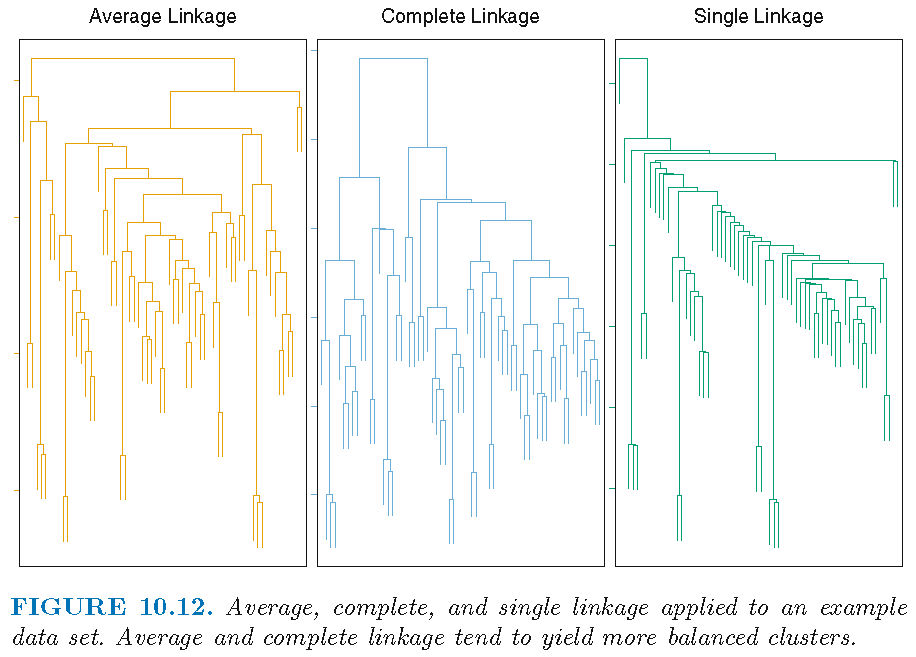
Hierarchical clustering (HC): practical issues
Overview on practical issues
Some practical issues on hierarchical clustering:
- What pairwise dissimilarity measure to use?
- Is standardization needed for observations for each feature?
- Is cluster found by a clustering algorithm sensible?
- How to interpret clustering results?
Choice of dissimilarity measure
The choice of dissimilarity measure
- is very important since it has a strong effect on the resulting dendrogram
- should aim at capturing latent patterns in the data
- should be determined by the type of data being clustered and the scientific question at hand
Correlation-based distance
- Correlation-based distance considers 2 observations to be similar if their features are highly correlated, even though the observed values may be far apart in terms of Euclidean distance.
- There are various measures on correlation, such as the (Pearson’s) sample correlation, Spearman’s rank correlation, Kendall’s rank correlation
- Distance based on Pearson’s sample correlation is also based on standardized entries in each observation, due to how this correlation is computed. Hence, such a distance is not effected by the magnitude of observations in terms of Euclidean distance
Correlation-based distance
- \(3\) observations \(\boldsymbol{x}_i = (x_{i1},\ldots,x_{i,10}), i=1,2,3\) for \(10\) features
- let \(r_{kl}\) be Pearson sample correlation between the entries of \(\boldsymbol{x}_k\) and the entries of \(\boldsymbol{x}_l\), i.e., \[ r_{kl} = \frac{1}{p-1}\sum_{j=1}^p \frac{(x_{kj}-\bar{x}_k)}{s_k}\frac{(x_{lj}-\bar{x}_l)}{s_l}, k \ne l \] with \(\bar{x}_i = p^{-1}\sum_{j=1}^p x_{ij}\) and \[s_i^2 = (p-1)^{-1}\sum_{j=1}^p (x_{ij}-\bar{x}_i)^2\]
- if \(\vert r_{12} \vert > \vert r_{13} \vert\), then \(\boldsymbol{x}_1\) is more similar to \(\boldsymbol{x}_2\) than it is to \(\boldsymbol{x}_3\)
Correlation-based distance
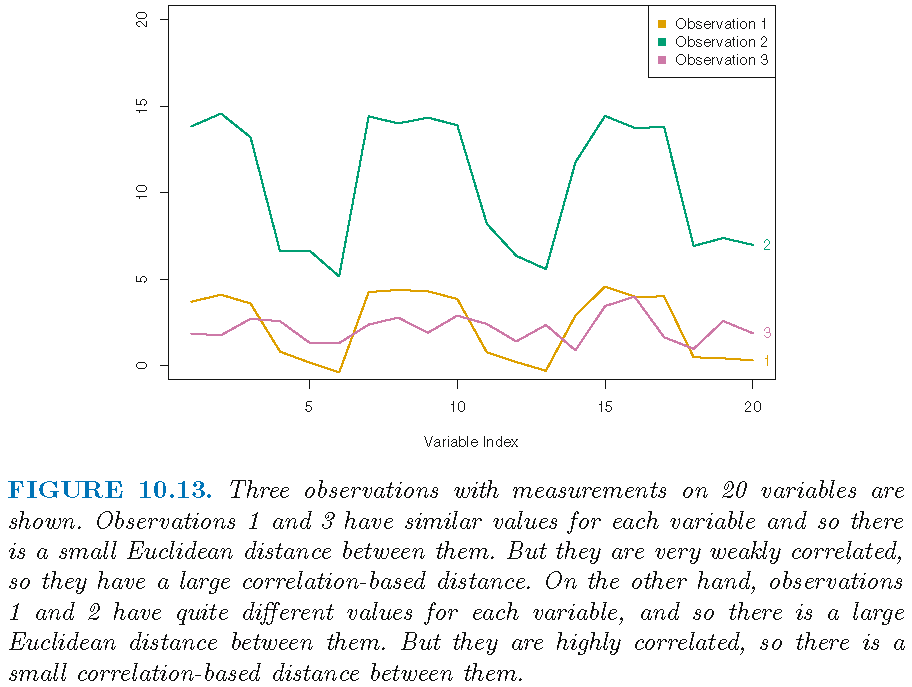
Choice of dissimilarity measure
Consider an online retailer interested in clustering shoppers based on their past shopping histories. The goal is to identify subgroups of similar shoppers, so that shoppers within each subgroup can be shown items and advertisements that are particularly likely to interest them.
Suppose the data takes the form of a matrix where the rows are the shoppers and the columns are the items available for purchase; the elements of the data matrix indicate the number of times a given shopper has purchased a given item (i.e. a 0 if the shopper has never purchased this item, a 1 if the shopper has purchased it once, etc.)
Choice of dissimilarity measure
What type of dissimilarity measure should be used to cluster the shoppers?
If Euclidean distance is used, then shoppers who have bought very few items overall (i.e. infrequent users of the online shopping site) will be clustered together. This may not be desirable.
If correlation-based distance is used, then shoppers with similar preferences (e.g. shoppers who have bought items A and B but never items C or D) will be clustered together, even if some shoppers with these preferences are higher-volume shoppers than others.
Standardization of observations
Consider an online retailer interested in clustering shoppers based on their past shopping histories. The goal is to identify subgroups of similar shoppers, so that shoppers within each subgroup can be shown items and advertisements that are particularly likely to interest them.
Suppose the data takes the form of a matrix where the rows are the shoppers and the columns are the items available for purchase; the elements of the data matrix indicate the number of times a given shopper has purchased a given item (i.e. a 0 if the shopper has never purchased this item, a 1 if the shopper has purchased it once, etc.)
Standardization of observations
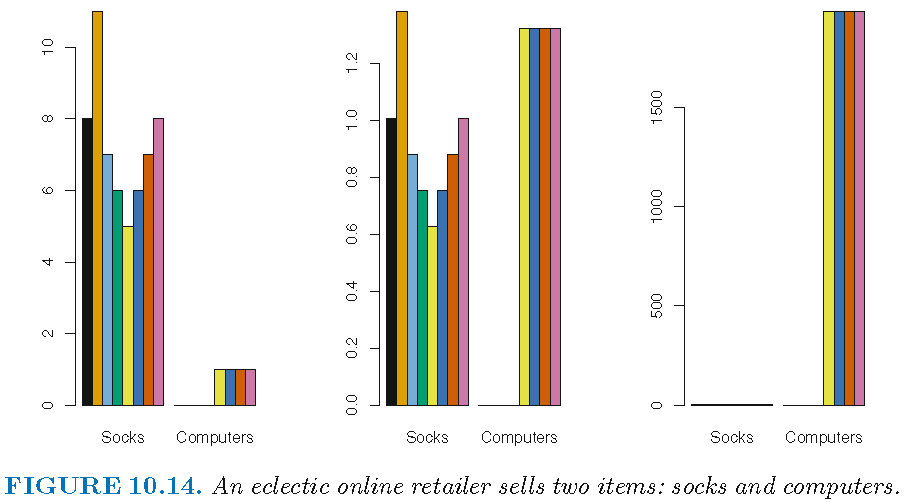
If we choose a dissimilarity measure that is easily affected the magnitudes of observations, then high-frequency purchases like socks tend to have a much larger effect on the inter-shopper dissimilarities, and hence on the clustering ultimately obtained, than rare purchases like computers. This may not be desirable.
If the variables are scaled to have standard deviation one before the inter-observation dissimilarities are computed, then each variable will in effect be given equal importance in the hierarchical clustering performed.
Standardization of observations
Left: raw data on purchase frequency; Center: scaled data on purchase frequency; Right: raw data on purchase expense
Standardization of observations
We might also want to scale the variables to have standard deviation one if they are measured on different scales; otherwise, the choice of units (e.g. centimeters versus kilometers) for a particular variable will greatly affect the dissimilarity measure obtained.
Whether or not it is a good decision to scale the variables before computing the dissimilarity measure depends on the application at hand.
The issue of whether or not to scale the variables before performing clustering applies to K-means clustering as well.
Validation of obtained clusters
- Clusters are always formed when clustering is performed. But do the clusters that have been found represent true subgroups in the data, or are they simply a result of clustering the noise?
- For instance, if we were to obtain an independent set of observations, then would those observations also display the same set of clusters? This is a hard question to answer.
- There exist a number of techniques for assigning a p-value to a cluster in order to assess whether there is more evidence for the cluster than one would expect due to chance. However, there has been no consensus on a single best approach.
Robustness on clustering
Suppose that most of the observations truly belong to a small number of (unknown) subgroups, and a small subset of the observations are quite different from each other and from all other observations.
Since both K-means and hierarchical clustering will assign each observation to a cluster, the clusters found may be heavily distorted due to the presence of outliers that do not belong to any cluster.
Mixture models are an attractive approach for accommodating the presence of such outliers.
Robustness on clustering
Suppose that we cluster \(n\) observations, and then cluster the observations again after removing a subset of the \(n\) observations at random.
We would hope that the two sets of clusters obtained would be quite similar. But often this is not the case!
Clustering methods generally are not very robust to perturbations to the data.
Recommendations on clustering
Perform clustering with different choices of standardization, dissimilarity measures, and linkages, and look at the full set of results in order to see what patterns consistently emerge.
Clustering subsets of the data in order to get a sense of the robustness of the clusters obtained.
Clustering results should not be taken as the absolute truth about a data set. Rather, they should constitute a starting point for the development of a scientific hypothesis and further study, preferably on an independent data set.
Hierarchical clustering (HC): software implementation
R software needed
R commands and libraries needed:
- R built-in command
distoras.distto compute pairwise dissimilarity - R built-in command
hclustapplies to pairwise dissimilarities to implement HC - R built-in command
cutreeapplies to the output ofhclustto obtain clusters - R library
ggdendroand commandggdendrogram{ggdendro}create dendrogram from the output ofhclust
Workflow
Workflow for software implementation:
- obtain pairwise dissimilarities
- implement hierarchical clustering
- show dendrogram
Command: dist
dist computes and returns the distance matrix that is obtained by using the specified distance measure to compute the distances between the rows of a data matrix. Its basic syntax is
dist(x, method = "euclidean")x: a numeric matrix, data frame or “dist” object.method: the distance measure to be used. It must be one of “euclidean”, “maximum”, “manhattan”, “canberra”, “binary” or “minkowski”.
Command: as.dist
as.dist converts an arbitrary square symmetric matrix into a form that hlust recognizes as a distance matrix. It can be used to induce correlation-based distance. Its basic syntax is
as.dist(x, ...)x: a numeric matrix, data frame or “dist” object.
Command: hclust
hclust implements hierarchical cluster analysis on a set of dissimilarities using a specified linkage. Its basic syntax is
hclust(d, method = "complete")d: a dissimilarity structure as produced bydistmethod: the agglomeration method to be used. It can be “single”, “complete”, “average” or “centroid”
Command: hclust
When clustering \(n\) observations, hclust returns:
height: a set of \(n-1\) real values that are called “clustering heights”. Each height is the value of the dissimilarity measure for the corresponding agglomeration.labels: labels for each of the objects being clustered.
Command: cutree
cutree cuts a tree, e.g., as resulting from hclust, into several groups either by specifying the desired number(s) of groups or the cut height(s). Its basic syntax is
cutree(tree, k = NULL, h = NULL)tree: a tree as produced byhclust.k: an integer scalar or vector with the desired number of groups.h: numeric scalar or vector with heights where the tree should be cut.
Command: ggdendrogram
ggdendrogram{ggdendro} creates a dendrogram using the output of hclust. Its basic syntax is
library(ggdendro)
ggdendrogram(data, leaf_labels = TRUE, rotate = FALSE)data: objects of classdendrogram,hclustortreeleaf_labels: ifTRUE, shows leaf labelsrotate: ifTRUE, rotates plot by 90 degrees
Note: the command plot can be applied to hclust object to create dendrogram
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19041)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] stringr_1.3.1 xfun_0.4 digest_0.6.18
[13] evaluate_0.12