Stat 437 Lecture Notes 4b
Xiongzhi Chen
Washington State University
Bayes classifier: practice
The two-group model
Prior probabilities for class labels: \[\Pr(Y_i =1)=\Pr(Y_i =2)=0.5\]
- Conditional density for \(x_i\) in Class 1: \[ f_1(x | Y_i=1) \sim \text{Gaussian}(\mu_1=(3,0),\mathbf{I}) \]
Conditional density for \(x_i\) in Class 2: \[ f_2(x | Y_i=2) \sim \text{Gaussian}(\mu_2=(0,-4),\mathbf{I}) \]
\(n=10\) observations \(x_i\)’s, which are independent given the \(n=10\) class labels \(Y_i\)
The two-group model
> set.seed(2)
> gp=rbinom(n=10, size=1, prob=0.5)
> gp1=which(gp==0); gp2=which(gp==1)
> x=matrix(rnorm(10*2), ncol=2)
> x[gp1,1]=x[gp1,1]+3; x[gp2,2]=x[gp2,2]-4gp=rbinom(...): generate 10 class labels randomly, \(0\) for Class 1 and \(1\) for Class 2gp1: indices of observations in Class 1gp2: indices of observations in Class 2x[gp1,]: observations from Class 1, with distribution \(\text{Gaussian}(\mu_1=(3,0),\mathbf{I})\)x[gp2,]: observations from Class 2, with distribution \(\text{Gaussian}(\mu_2=(0,-4),\mathbf{I})\)
Generated observations
> colnames(x)=c("V1","V2")
> y = as.data.frame(x); y$obId = 1:10; y$Class=as.factor(gp+1)
> y
V1 V2 obId Class
1 3.1324203 -2.311069 1 1
2 0.7079547 -3.121395 2 2
3 -0.2396980 -3.964193 3 2
4 4.9844739 1.012829 4 1
5 -0.1387870 -3.567735 5 2
6 0.4176508 -1.909181 6 2
7 3.9817528 -1.199926 7 1
8 -0.3926954 -2.410362 8 2
9 1.9603310 1.954652 9 1
10 1.7822290 -3.995062 10 2obId: observation id- observation \(x_i\) is in row \(i\), with coordinates
V1andV2 Class: class label for each \(x_i\)
Visualize data
> library(ggplot2)
> ggplot(y,aes(V1,V2))+theme_bw()+guides(color = FALSE)+
+ geom_label(aes(label=obId,color=Class))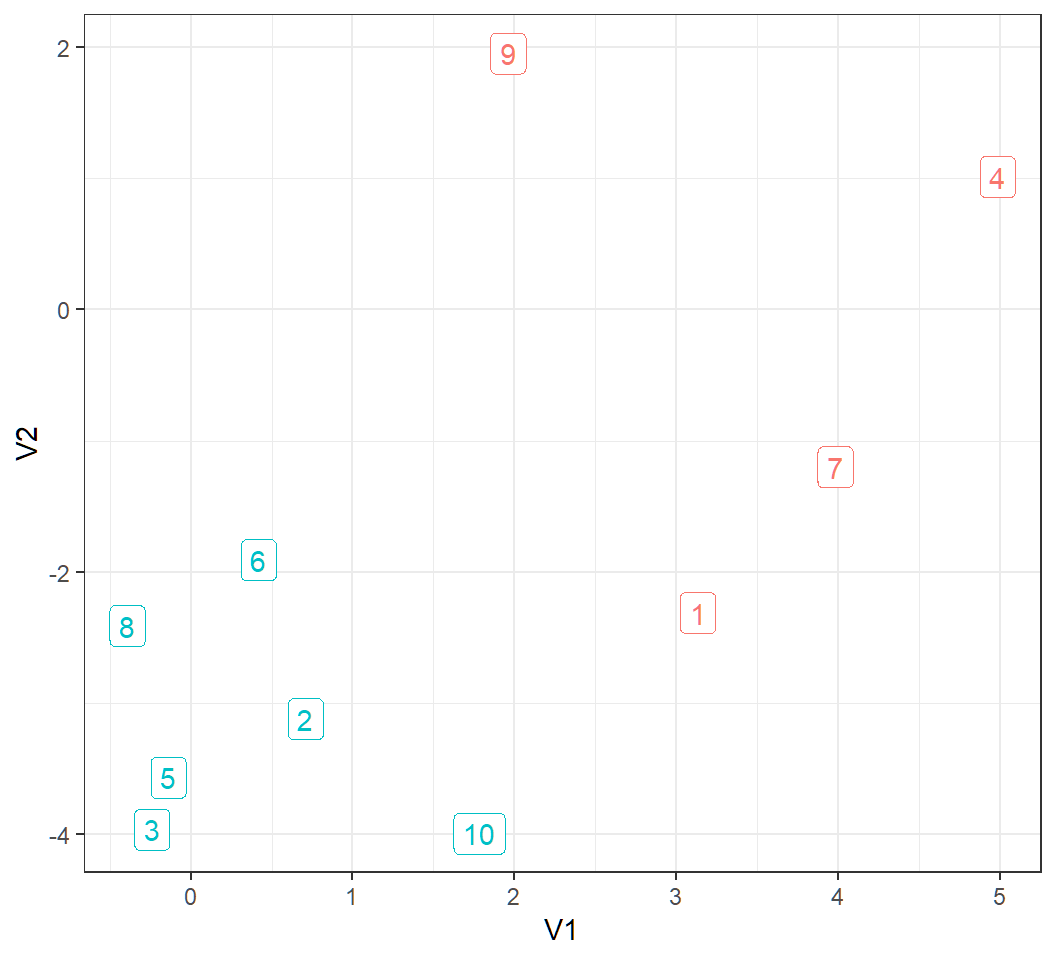
Marginal and posterior distr.
marginal density of \(x_i\) is \[ \begin{aligned} f(x) & = f_1(x | Y_i=1) \Pr(Y_i =1) + f_2(x | Y_i=2) \Pr(Y_i =2)\\ & = 0.5f_1(x | Y_i=1) + 0.5 f_2(x | Y_i=2) \end{aligned} \]
- posterior probability of \(x_i\) belonging to Class \(j\) is \[ \begin{aligned} \Pr(Y_i=j|x_i) & = \frac{f_j(x_i | Y_i=j) \Pr(Y_i =j)}{f(x_i)} \\ & = \frac{0.5 f_j(x_i | Y_i=j)}{f(x_i)} \end{aligned} \]
need \(f_j(x_i | Y_i=j)\) to get marginal and posterior
Obtain posterior probability
> library(mvtnorm) # compute multivariate Gaussian probabilities
> postPClass = matrix(0,ncol=2,nrow=10) # store prosterior probabilities
> for (i in 1:10) {
+ cndp1 = dmvnorm(x[i,], mean =c(3,0), sigma = diag(2))
+ cndp2 = dmvnorm(x[i,], mean =c(0,-4), sigma = diag(2))
+ fmarginal = 0.5*cndp1 +0.5*cndp2
+ jp1=0.5*cndp1; jp2=0.5*cndp2
+ postp1=jp1/fmarginal; postp2=jp2/fmarginal
+ postPClass[i,]=c(postp1,postp2)
+ }
> colnames(postPClass)=c("PPCL1","PPCL2")
> postPClassdmvnorm(x[i,], mean =c(3,0), sigma = diag(2)): \(f_1(x_i | Y_i=1) \sim \text{Gaussian}(\mu_1=(3,0),\mathbf{I})\)fmarginal: \(f(x_i)\), andjp1: \(f_1(x_i | Y_i=1) \Pr(Y_i =1)\)postp1andpostp2: \(\Pr(Y_i=1|x_i)\) and \(\Pr(Y_i=2|x_i)\)
Posterior probabilities
PPCL1 PPCL2
[1,] 9.747409e-01 2.525912e-02
[2,] 1.046019e-03 9.989540e-01
[3,] 2.095184e-06 9.999979e-01
[4,] 1.000000e+00 1.683903e-10
[5,] 1.384877e-05 9.999862e-01
[6,] 5.296168e-02 9.470383e-01
[7,] 9.999762e-01 2.380586e-05
[8,] 6.620328e-04 9.993380e-01
[9,] 1.000000e+00 3.390852e-08
[10,] 7.972362e-04 9.992028e-01- column 1
PPCL1: posterior probability of an observation belonging to Class 1 - column 2
PPCL2: posterior probability of an observation belonging to Class 2
Bayes classifier
Bayes classifier assigns \(x_i\) to Class \(j\) if \(\Pr(Y_i=j|x_i)\) is the larger among the two \(\Pr(Y_i=k|x_i)\) for \(k=1\) or \(2\):
> postPClass=data.frame(postPClass)
> postPClass$TrueClass = rep(1,10)
> postPClass$TrueClass[gp2]=2 #true class labels
> postPClass$EstClass=rep(1,10)
> # compare posterior probabilities
> comp = (postPClass$PPCL1 >= postPClass$PPCL2)
> postPClass$EstClass[comp==TRUE]=1 # assign Class 1
> postPClass$EstClass[comp==FALSE]=2 # assign Class 2
> postPClasspostPClass$PPCL1 >= postPClass$PPCL2: check if \(\Pr(Y_i=1|x_i)>\Pr(Y_i=2|x_i)\) for each \(i\); return alogicalvector
Bayes classifier
PPCL1 PPCL2 TrueClass EstClass
1 9.747409e-01 2.525912e-02 1 1
2 1.046019e-03 9.989540e-01 2 2
3 2.095184e-06 9.999979e-01 2 2
4 1.000000e+00 1.683903e-10 1 1
5 1.384877e-05 9.999862e-01 2 2
6 5.296168e-02 9.470383e-01 2 2
7 9.999762e-01 2.380586e-05 1 1
8 6.620328e-04 9.993380e-01 2 2
9 1.000000e+00 3.390852e-08 1 1
10 7.972362e-04 9.992028e-01 2 2TrueClass: true class labelsEstClass: labels obtained by the Bayes classifier
Classification error
- \(\hat{y}_i\): estimated class label for true class label \(y_i\) for \(x_i\)
- \(L\): 0-1 loss, i.e., \(L(y_i,\hat{y}_i)=0\) if \(y_i=\hat{y}_i\), and \(1\) otherwise
Classification error, defined as \[ e_n = \frac{1}{n} \sum_{i=1}^n L(y_i,\hat{y}_i), \] is zero for this example:
> table(EstClass,TrueClass)
TrueClass
EstClass 1 2
1 4 0
2 0 6k-nearest neighbor (kNN) classifiers: software implementation
R software needed and workflow
R commands and libraries needed:
- R library
classand commandknn{class}for kNN classification
General workflow for kNN classifiers:
- standardize each feature (if measurements on different features are on quite different scales since the Euclidean distance is used as a measure on how close two observations are in the construction of neighborhoods)
- choose an optimal neighborhood size \(k\) (e.g., by cross-validation)
- apply optimal kNN classifier
Command knn{class}
knn{class} implements kNN classification for test set from training set:
- for each row of the test set, the
knearest vectors (in Euclidean distance) from the training set are found and selected - the classification is decided by majority vote, based on the class labels of the
knearest vectors - if there are ties (in Euclidean distance) for the
kth nearest vector among the selected vectors, then all such “tied” vectors are included in the vote
Command knn{class}
Its basic syntax is
knn(train, test, cl, k = 1, prob = FALSE)train: matrix or data frame of training set casestest: matrix or data frame of test set cases. A vector will be interpreted as a row vector for a single casecl: factor of true classifications of training setk: number of neighbors to useprob: if set astrue, the proportion of the votes for the winning class is returned as attributeprob
k-nearest neighbor (kNN) classifiers: Example 1
Generate data
- Randomly generate 30 independent observations from a tri-variate Gaussian distribution with mean 0 and identity covariance matrix
> set.seed(1); x=matrix(rnorm(30*3), ncol=3)
> colnames(x) = c("X1","X2","X3"); x = as.data.frame(x)
> x$class= rep(1,nrow(x))xhas 30 rows and 3 columns, i.e., there are 30 observations for 3 featuresthe 30 observations form 1 class (judged from the distribution they follow)
Visualize data
> library(ggplot2)
> ggplot(data=x,aes(X1,X2,color=X3))+geom_point()+theme_bw()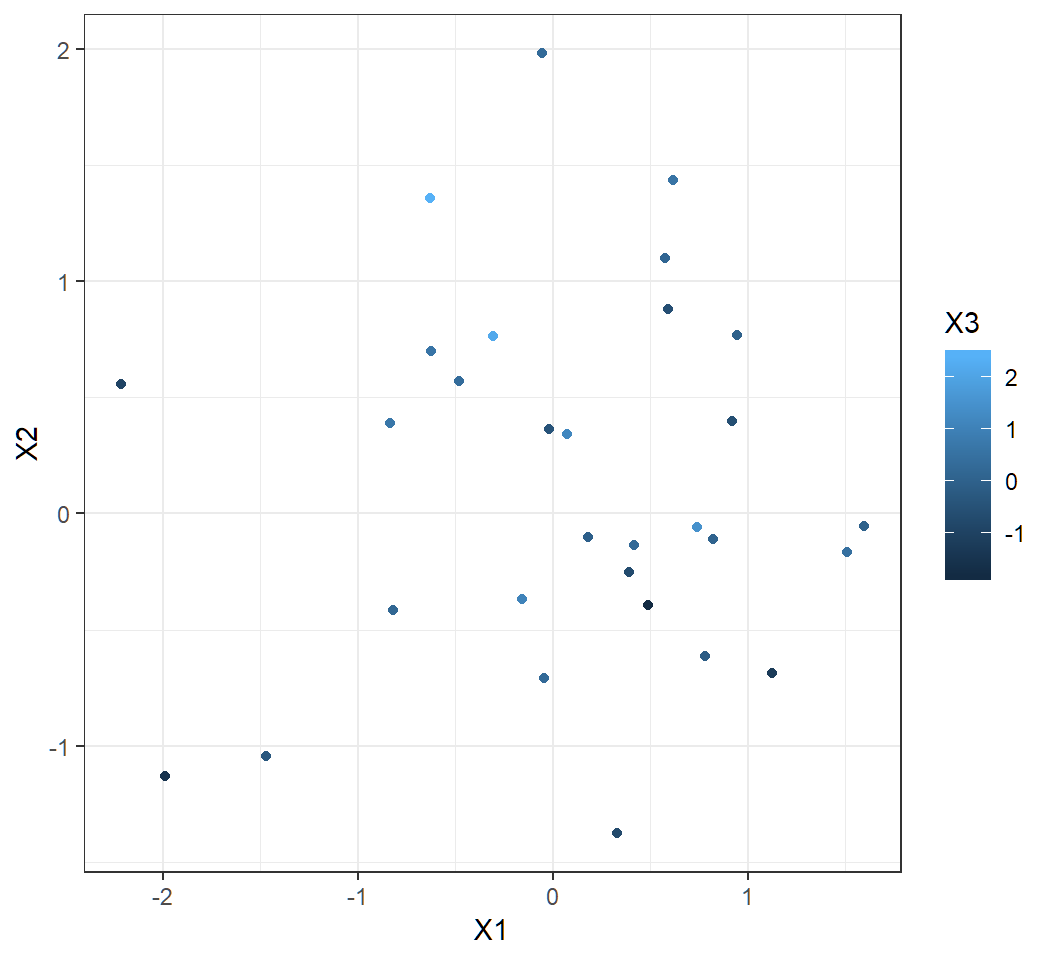
Split data
- set the first 20 observations in
xas training set - set the rest 10 observations in
xas test set
> trainingSet =x[1:20,1:3]; trainingLabels =x[1:20,4]
> testSet =x[21:30,1:3]; testLabels =x[21:30,4]Note: in terms of software implementation via knn, training set and test set contain observations for features but not class labels
kNN: k=1 or 3
Fact: \(k\)NN classification error is \(0\) for all \(k\) when there is only one class (since the average label is always the exact, true class label)
> library(class)
> knn1 = knn(trainingSet,testSet, cl=trainingLabels, k = 1)
> table(knn1,testLabels)
testLabels
knn1 1
1 10
> knn3 = knn(trainingSet,testSet, cl=trainingLabels, k = 3)
> table(knn3,testLabels)
testLabels
knn3 1
1 10- 0 classification error
kNN: k=10 or 30
Fact: \(k\)NN classification error is \(0\) for all \(k\) when there is only one class (since the average label is always the exact, true class label)
> library(class)
> knn10 = knn(trainingSet,testSet, cl=trainingLabels, k = 10)
> table(knn10,testLabels)
testLabels
knn10 1
1 10
> knn30 = knn(trainingSet,testSet, cl=trainingLabels, k = 30)
> table(knn30,testLabels)
testLabels
knn30 1
1 10- 0 classification error
kNN: k bigger than sample size
kNN classifier with \(k > n\), i.e., neighborhood size bigger than sample size; \(n=30\), \(k=31\), \(k=40\):
> library(class)
> knn(trainingSet,testSet, cl=trainingLabels, k = 31)
Warning in knn(trainingSet, testSet, cl = trainingLabels, k =
31): k = 31 exceeds number 20 of patterns
[1] 1 1 1 1 1 1 1 1 1 1
Levels: 1
> knn(trainingSet,testSet, cl=trainingLabels, k = 40)
Warning in knn(trainingSet, testSet, cl = trainingLabels, k =
40): k = 40 exceeds number 20 of patterns
[1] 1 1 1 1 1 1 1 1 1 1
Levels: 1- 0 classification error but with warnings
- insensible to pick \(k >n_1\), where \(n_1\) is the sample size for training set
k-nearest neighbor (kNN) classifiers: Example 2
Human cancer microarray data
Human cancer microarray data:
- 6830 genes (i.e., 6830 features)
- 64 samples (i.e., 64 cancer cell lines)
- 14 different cancer types; information on cancer types at wiki-NCI60
The data matrix has dimension \(6830 \times 64\), each of whose columns is an observation for all features
We will pick 3 caner types “BREAST”, “LEUKEMIA”, “COLON”, and randomly select the same set of \(50\) genes for each cancer type, to balance computations and illustrate kNN classifiers
Obtain subset of data
> library(ElemStatLearn) # library containing data
> data(nci); n = dim(nci)[2]; p = dim(nci)[1] #get dimensions
> set.seed(123)
> rSel = sample(1:p, size=50, replace = FALSE)
> chk = colnames(nci) %in% c("BREAST", "LEUKEMIA", "COLON")
> cSel = which(chk ==TRUE)
> ncia = nci[rSel,cSel]
> colnames(ncia) = colnames(nci)[cSel]colnames(nci): each column name is the cancer type for an observationchk: check if a cancer type is “BREAST”, “LEUKEMIA” or “COLON”- take features (i.e., genes) from rows of
nci, and take samples (i.e., observations for features) from columns ofnci
Glimpse on subset of data
First few entries of the transposed subset data:
> ncib=data.frame(t(ncia)); ncib$Class=colnames(ncia)
> dim(ncib)
[1] 20 51
> head(ncib[1:3,1:5])
X1 X2 X3 X4 X5
BREAST -0.075 1.1725 0.510 0.385 0.765
BREAST.1 -0.570 0.0975 0.565 -0.830 2.130
BREAST.2 0.000 1.0375 1.175 -0.160 -0.150For ncib:
- 20 rows; 1st 50 entries of each row form an observation for all 50 features
- last entry of each row is the cancer type of an observation
Visualize subset of data
> library(ggplot2)
> ggplot(ncib,aes(X1,X2,color=Class))+geom_point()+theme_bw()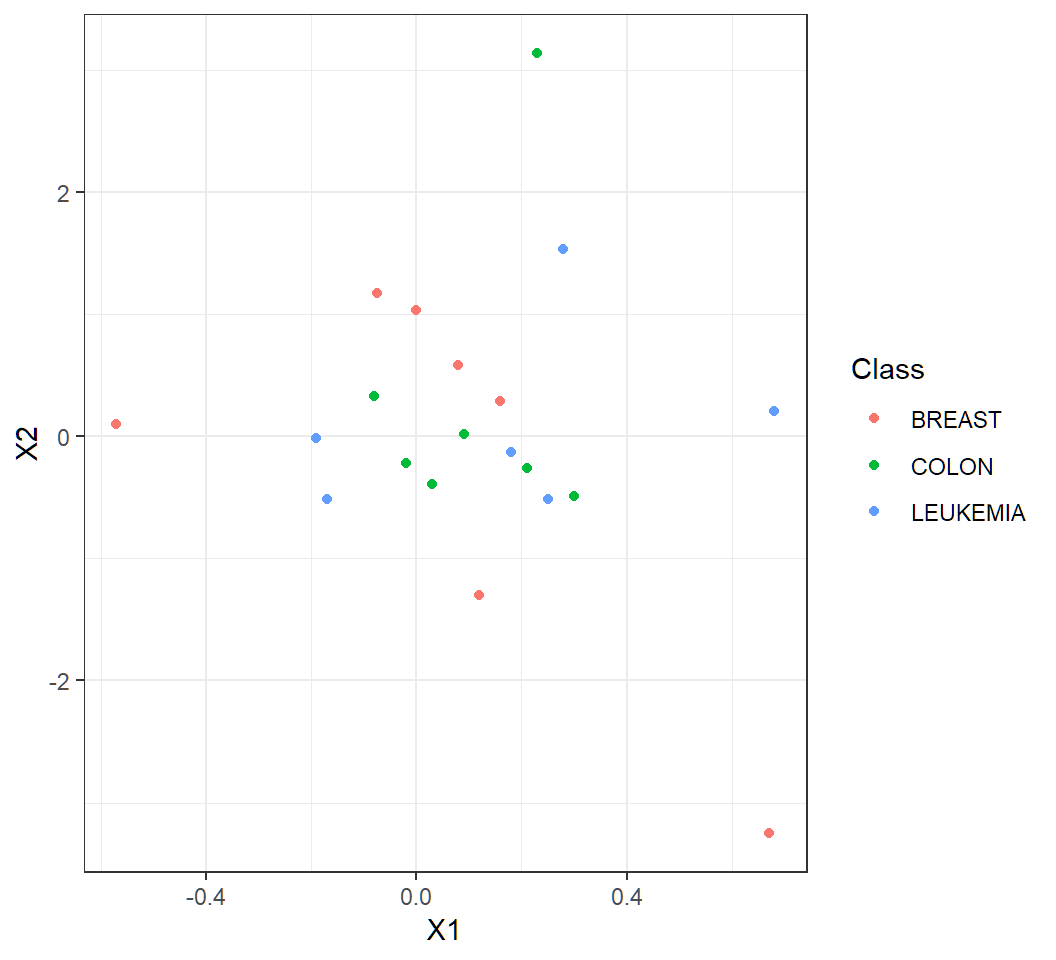
Standardize and split data
60% of standardized data as training set, and the rest 40% as test set:
> ncibsd=scale(ncib[,1:50]); set.seed(1)
> rTrain= base::sample(1:nrow(ncibsd),0.6*nrow(ncibsd))
> rTest =(1:nrow(ncibsd))[-rTrain]
> trainSet =ncibsd[rTrain,]; testSet =ncibsd[rTest,]
> trainLabels=ncib$Class[rTrain]; testLabels=ncib$Class[rTest]scale(ncib[,1:50]): standardize each feature (i.e., each column) of data- last column of
ncibcontains class labels
kNN: k=2
> library(class)
> knn2eg2= knn(trainSet,testSet, cl=trainLabels, k = 2)
> length(testLabels)
[1] 8
> testLabels
[1] "LEUKEMIA" "LEUKEMIA" "COLON" "COLON" "COLON"
[6] "BREAST" "BREAST" "BREAST"
> table(knn2eg2,testLabels)
testLabels
knn2eg2 BREAST COLON LEUKEMIA
BREAST 1 0 0
COLON 2 3 0
LEUKEMIA 0 0 2
> # classification error
> sum(1- as.numeric(knn2eg2==testLabels))/length(testLabels)
[1] 0.25- \(8\) test observations; \(2\) of them are misclassified (i.e., \(2\) observations for “BREAST” cancer are misclassified)
- classification error is \(2/8=0.25\)
kNN: k=4
> library(class)
> knn4eg2= knn(trainSet,testSet, cl=trainLabels, k = 4)
> length(testLabels)
[1] 8
> testLabels
[1] "LEUKEMIA" "LEUKEMIA" "COLON" "COLON" "COLON"
[6] "BREAST" "BREAST" "BREAST"
> table(knn4eg2,testLabels)
testLabels
knn4eg2 BREAST COLON LEUKEMIA
BREAST 1 0 0
COLON 1 2 0
LEUKEMIA 1 1 2
> # classification error
> sum(1- as.numeric(knn4eg2==testLabels))/length(testLabels)
[1] 0.375- \(8\) test observations; \(3\) of them are misclassified (i.e., \(2\) observations for “BREAST” cancer and \(1\) observation for “COLON” cancer are misclassified)
- classification error is \(3/8=0.375\)
Remarks
For the example, the data set we investigated has only \(20\) observations. So, it is not very sensible to try cross-validation to choose an optimal neighborhood size \(k\)
k-nearest neighbor (kNN) classifiers: Example 3
Iris flower data
- For each of 3 species “setosa”, “versicolor”, “virginica” of irises, 50 observations on 4 features
Sepal.Length,Sepal.Width,Petal.LengthandPetal.Widthare stored in objectiris(in R libraryggplot2orclass)
> library(class); dim(iris)
[1] 150 5
> iris[1,]
Sepal.Length Sepal.Width Petal.Length Petal.Width Species
1 5.1 3.5 1.4 0.2 setosa
> unique(iris$Species)
[1] setosa versicolor virginica
Levels: setosa versicolor virginica- The target is to classify an iris plant into one of the three species using these measurements
Data splitting and cross-validation
Proposal for data splitting and cross-validation (cv):
- 80% of data as training set that contains 40 random samples for each species, and the rest 20% as test set
- apply “\(m\)-fold cross-validation” on the training set to choose optimal \(k\) for kNN classifiers
- apply optimal kNN classifier to test set
Note: different proportions can be used for data splitting, and they should ensure that a learning method be well trained; usually 5-fold or 10-fold cross-validation is used
Estimating test error
- We cannot directly estimate test error using test set since we do not know the class labels for observations in this set
- So, we split the training set into 2 disjoint subsets: one as “training set” and the other as “test set”, for which we know the class labels for their observations, and test error is estimated using such “test set”; namely, part of the training set is used as a test set
- \(m\)-fold cross-validation implements the above strategy by randomly splitting the training set into \(m\) folds of approximately equal size, uses 1 fold as “test set” and the rest as “training set”, and does this for each of the \(m\) folds; see Chapter 5 of Text for details on cross-validation
Splitting data: Step 0
> library(class); set.seed(314) # seed needed!!
> trainId=c(sample(1:50,40),sample(51:100,40),
+ sample(101:150,40))
> testId = (1:150)[-trainId]
> trainingSet=iris[trainId,1:4]; testSet=iris[testId,1:4]
> trainingLabs=iris$Species[trainId]
> testLabs=iris$Species[testId]trainId=c(...): observation id’s for training set; in data matrixiris, 1st 50 rows (i.e., observations) are for “setosa”, 2nd 50 for “versicolor”, and 3rd 50 for “virginica”trainingSetcontains 40 random samples for each species;testSetis the test settrainingLabs(ortestLabs): class labels for observations in training set (or tet set)
Cross-validation: Step 1
\(10\)-fold random split of training set:
> m=10; set.seed(123) # seed needed!!
> nrow(trainingSet)
[1] 120
> folds=sample(1:m,nrow(trainingSet),replace=TRUE)
> folds[1:10]
[1] 3 8 5 9 10 1 6 9 6 5folds=sample(...,replace=TRUE): sample with replacement (i.e.,replace=TRUE) entries of the vector \((1,2,\ldots,m)\)120=nrow(trainingSet)times, and store sampled entries intofoldsfoldshas120=nrow(trainingSet)entries, and the entries are numbers between \(1\) and \(m=10\) and can be repeated
Cross-validation: Step 1 (cont’d)
> which(folds==1) # obs. id's in fold 1
[1] 6 18 35 51 62 74 98 113
> table(folds)
folds
1 2 3 4 5 6 7 8 9 10
8 14 11 11 18 10 12 12 12 12 - for each
sbetween 1 and \(m\),which(folds==s)returns the id’s of observations in folds table(folds)shows, in the 2nd row of its output, the number of observations in each fold (with fold number stored infolds), which equivalently checks if folds have approximately equal size
Cross-validation: Step 2
\(10\)-fold cv for 2-NN classifier on training Set:
> k=2 # k for kNN; m=10
> testError1 = double(m) # store test error for each fold s
> for (s in 1:m) { # loop through s=1,...,m
+ trainingTmp =trainingSet[folds !=s,]
+ testTmp =trainingSet[folds==s,]
+ trainingLabsTmp =trainingLabs[folds !=s]
+ testLabsTmp =trainingLabs[folds==s]
+ knn2= knn(trainingTmp,testTmp,trainingLabsTmp,k)
+ nOfMissObs= sum(1-as.numeric(knn2==testLabsTmp))
+ terror=nOfMissObs/length(testLabsTmp) # test error
+ testError1[s]=terror
+ } # end of loop- for each
s, folds, i.e.,testTmp, is “test set” with class labelstestLabsTmp, and other folds, i.e.,trainingTmp, is “training set” with class labelstrainingLabsTmp
Cross-validation: Step 2 (cont’d)
> mean(testError1)
[1] 0.05040404
> sd(testError1)
[1] 0.04484363testError1: a vector of \(m\) entries, whose \(s\)th entry is the estimated test error obtained by setting foldsas test set, asschanges from 1 to \(m\)- the sample mean of all \(m\) entries of
testError1is regarded as the estimated test error for the given kNN classifier (k=2 in the example codes)
Cross-validation: summary
Summary for \(m\)-fold cross-validation for a kNN classifier:
- Randomly split \(n\) observations into \(m\) folds of approximately equal size
- Pick fold \(s\) as “test set”, set the remaining folds as “training set”, apply the kNN classifier, and obtain test error \(e_{s}\)
- Do Item 2. for each \(s\) in \(\{1,\ldots,m\}\), and obtain \(m\) \(e_{s}\)’s
- Compute sample mean \(\hat{\mu}(k,m)\) and sample standard deviation \(\hat{\sigma}(k,m)\) for the \(m\) \(e_{s}\)’s
- Use \(\hat{\mu}(k,m)\) as an estimate of the test error of the classifier
Note: we call \(e_{s}\) “test error” even though it is an estimate of test error
Cross-validation: Step 3
Choose optimal \(k\) via \(m\)-fold cross-validation for kNN classifiers using training set:
- Pick a sequence \(C\) of \(b\) distinct values for \(k\)
- Do Step 1 and Step 2 for each \(k\) in \(C\), obtain \(b\) estimated test errors, \(\hat{\mu}(k,m), k \in C\), for the \(b\) kNN classifiers
- Set as the optimal \(\hat{k}\) for which \(\hat{\mu}(\hat{k},m)\) is the smallest among \(\hat{\mu}(k,m), k \in C\), i.e., \[\hat{k} = \operatorname*{argmin}\{k \in C: \hat{\mu}(k,m)\}\]
Note: if we have enough computational power, we can pick \(C=\{1,2,\ldots,k_{\max}\}\) for a large number \(k_{\max}\)
Cross-validation: Step 3 (cont’d)
- If there are multiple optimal \(\hat{k}\)’s, then set the optimal as the one whose corresponding estimated test error has the smallest sample standard deviation for these optimal kNN classifiers
- For example, if \(\hat{\mu}(\hat{k}_1,m)\) and \(\hat{\mu}(\hat{k}_2,m)\) are both the smallest among \(\hat{\mu}(k,m), k \in C\) but \(\hat{\sigma}(\hat{k}_1,m) < \hat{\sigma}(\hat{k}_2,m)\), then pick \(\hat{k}_1\) as the optimal
- If further the estimated test errors of the kNN classifiers with these optimal \(\hat{k}\)’s have the same sample standard deviation, then pick the optimal as the smallest among these \(\hat{k}\) (which can reduce the computational cost of building neighborhoods for the optimal kNN classifier)
Cross-validation: Step 3
\(10\)-fold cv for kNN classifiers for \(k=1,\ldots,20\):
> kmax=20 # m=10 fold cv
> testErrors = matrix(0,nrow=2,ncol=kmax)
> for (k in 1:kmax) { # loop through k
+ testError1 = double(m) # store test errors for each k
+ for (s in 1:m) { # loop through s
+ trainingTmp =trainingSet[folds !=s,]
+ testTmp =trainingSet[folds==s,]
+ trainingLabsTmp =trainingLabs[folds !=s]
+ testLabsTmp =trainingLabs[folds==s]
+ knntmp= knn(trainingTmp,testTmp,trainingLabsTmp,k)
+ nOfMissObs= sum(1-as.numeric(knntmp==testLabsTmp))
+ terror=nOfMissObs/length(testLabsTmp) # test error
+ testError1[s]=terror } # loop in s ends
+ testErrors[,k]=c(mean(testError1),sd(testError1))
+ } # loop in k endstestErrors: each column contains the sample mean and sample standard deviation for the estimated test errors for a kNN classifier as \(k\) varies from \(1\) tokmax
Cross-validation: Step 3 (cont’d)
Optimal \(\hat{k}\) chosen by \(10\)-fold cv for \(k=1,\ldots,20\):
> colnames(testErrors)= paste("k=",1:kmax,sep="")
> rownames(testErrors)=c("mean(TestError)","sd(TestError)")
> testErrors=as.data.frame(testErrors)
> as.numeric(testErrors[1,])
[1] 0.04088023 0.04643579 0.04802309 0.03337662 0.03337662
[6] 0.03893218 0.03456710 0.04012266 0.03178932 0.03532468
[11] 0.04088023 0.04088023 0.04921356 0.04365801 0.05476912
[16] 0.04088023 0.04088023 0.04921356 0.05476912 0.04088023
> hatk=which(testErrors[1,]==min(testErrors[1,]))
> hatk
[1] 9hatk=which(...): find optimal \(\hat{k}\) for which the sample mean of the estimated test error is the smallest as \(k\) varies from \(1\) tokmax; the optimal \(\hat{k}=9\)
Cross-validation: extra
If we want to do \(m\)-fold cross-validation for kNN classifiers with \(k\) values in a sequence \(C\) using a different \(m\)-fold split of the training set, we only need to
Do Step 0 with a different
set.seedvalue to get differentfoldsvia the commandsample, and thenDo Step 1, 2 and 3
Apply optimal kNN classifier
Apply optimal classifier to test set:
> library(class)
> knnOpt= knn(trainingSet,testSet,trainingLabs,hatk)
> nOfMissObs1= sum(1-as.numeric(knnOpt==testLabs))
> terrorOpt=nOfMissObs1/length(testLabs) # test error
> terrorOpt
[1] 0
> table(knnOpt,testLabs)
testLabs
knnOpt setosa versicolor virginica
setosa 10 0 0
versicolor 0 10 0
virginica 0 0 10hatk: the optimal \(\hat{k}=9\) obtained by \(10\)-fold cross-validation on training settrainingSet- original training set
trainingSetis used by the optimal \(9\)-NN classifier to classify observations intestSet
Visualization via 2 features
> testSet$Species=testLabs
> testSet$EstimatedSpecies=knnOpt
> library(ggplot2)
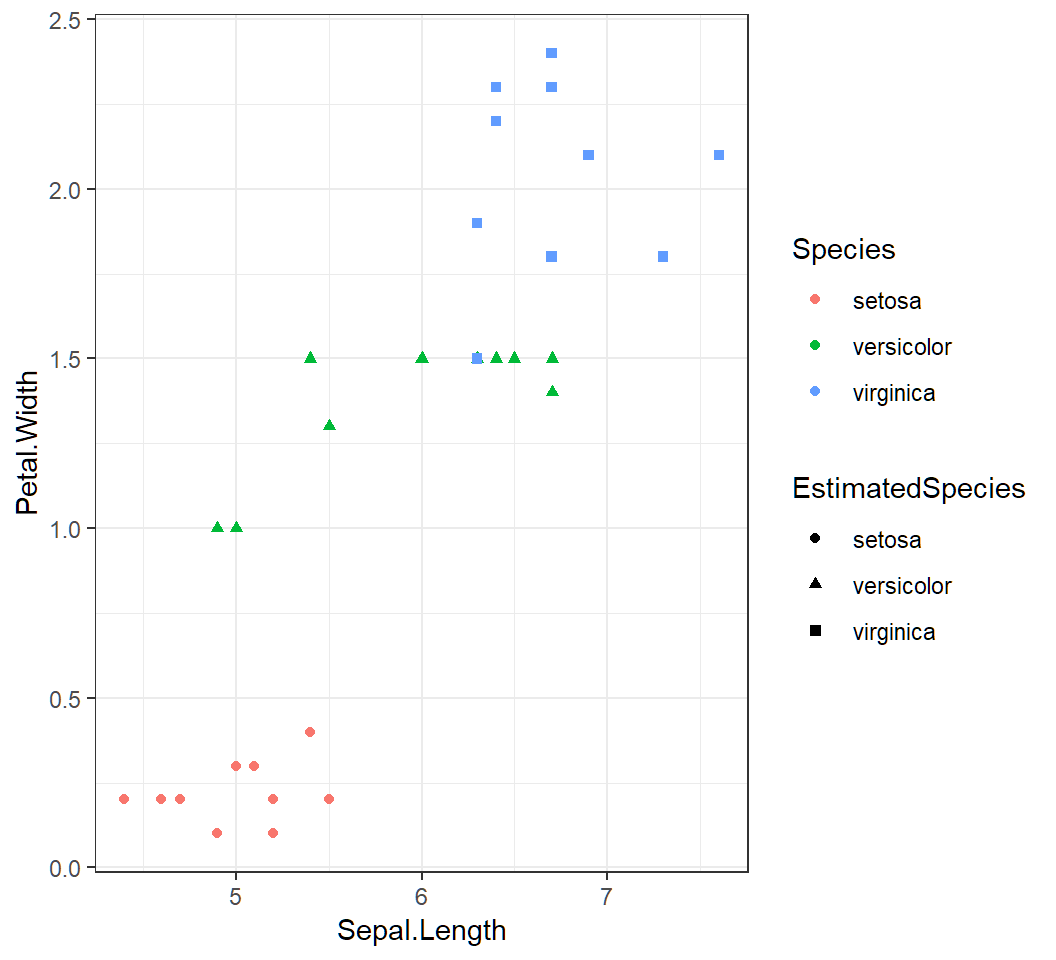
> cp =ggplot(testSet,aes(Sepal.Length,Petal.Width))+
+ geom_point(aes(shape=EstimatedSpecies,color=Species))+
+ theme_bw()- add to
testSetthe trueSpecieslabels and estimated species labelsEstimatedSpecies - create plot by
ggplotusing featuresSepal.LengthandPetal.Widthwithshapeandcoloraesthetics
Visualization via 2 features
Perfect classification:
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19041)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] stringr_1.3.1 xfun_0.4 digest_0.6.18
[13] evaluate_0.12