Stat 437 Lecture Notes 5c
Xiongzhi Chen
Washington State University
Discriminant analysis: Example 3
Human cancer microarray data
Human cancer microarray data:
- 6830 genes (i.e., 6830 features)
- 64 samples (i.e., 64 cancer cell lines)
- 14 different cancer types; information on cancer types at wiki-NCI60
The data matrix has dimension \(6830 \times 64\), each of whose columns is an observation for all features.
We will pick 3 caner types “BREAST”, “LEUKEMIA” and “COLON”, and randomly select the same set of \(50\) genes for each cancer type
Obtain subset of data
> library(ElemStatLearn) # library containing data
> data(nci); n0=dim(nci)[2]; p0=dim(nci)[1] #get dimensions
> set.seed(123)
> rSel = sample(1:p0, size=50, replace = FALSE)
> chk = colnames(nci) %in% c("BREAST", "LEUKEMIA", "COLON")
> cSel = which(chk ==TRUE)
> ncia = nci[rSel,cSel]
> colnames(ncia) = colnames(nci)[cSel]colnames(nci): each column name is the cancer type for an observationchk: check if a cancer type is “BREAST”, “LEUKEMIA” or “COLON”- take features (i.e., genes) from rows of
nci, and take samples (i.e., observations for features) from columns ofnci
Glimpse on subset of data
> # transposed subset data as `tmp`
> tmp = data.frame(t(ncia)); tmp$Class=colnames(ncia)
> rownames(tmp)=NULL; dim(tmp)
[1] 20 51
> table(tmp$Class)
BREAST COLON LEUKEMIA
7 7 6
> tmp[1:3,(ncol(tmp)-3):ncol(tmp)]
X48 X49 X50 Class
1 -0.415 -0.735 1.935 BREAST
2 -0.040 -0.780 0.060 BREAST
3 0.750 -0.320 1.800 BREAST- the last entry of each row of
tmpis the cancer type of an observation, and the rest in each such row is an observation on the features - \(p=50\) feature variables \(X_1,\ldots,X_{50}\), and \(n=20\) observations; \(p > 2n\)
Visualize subset of data
In terms of class memberships, observations overlap each other much?
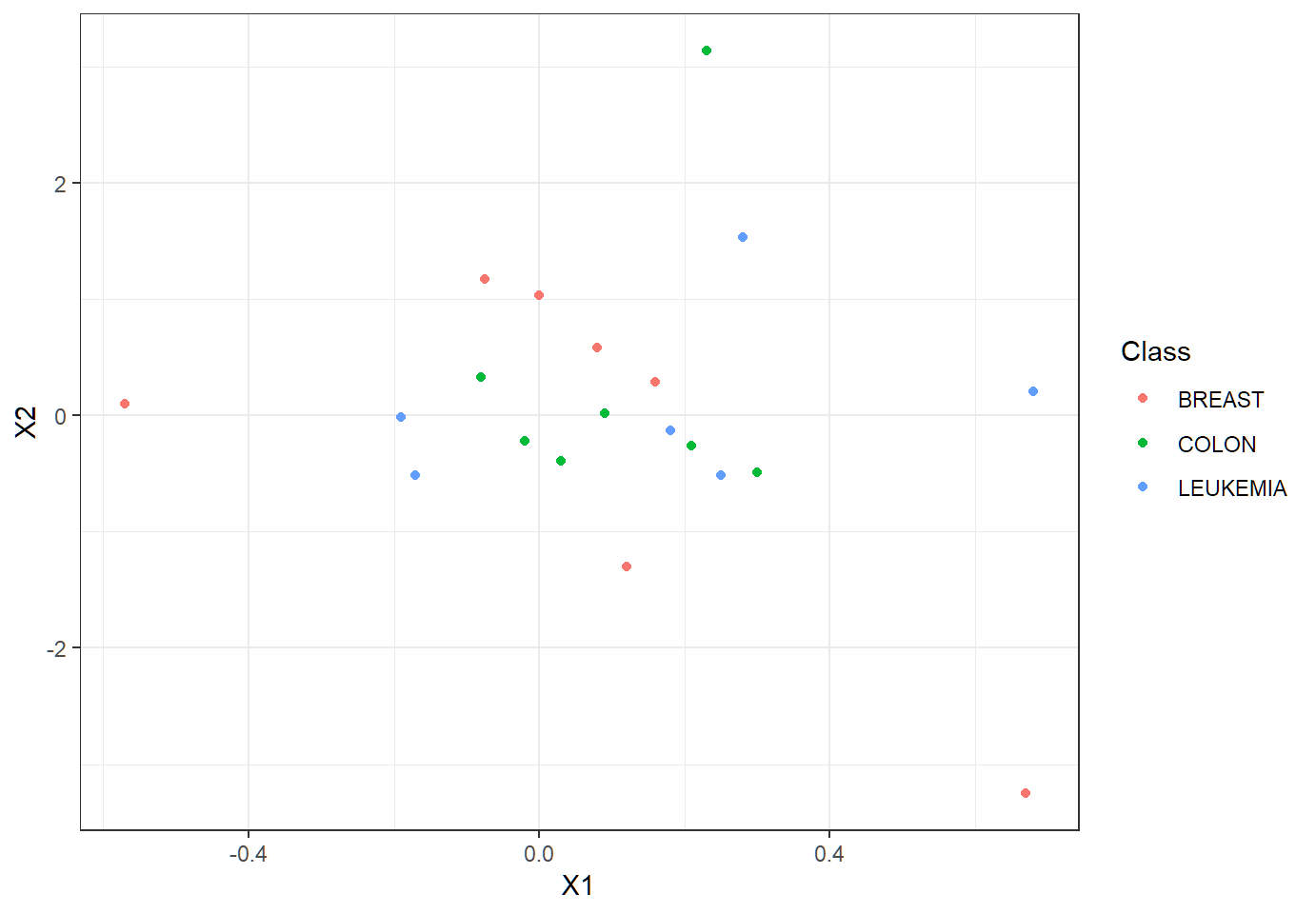
Visualize subset of data
Each observation on the \(p=50\) features (is in \(\mathbb{R}^{50}\) and) is a 50-dimensional vector.
- When such observations are visualized via only 2 features, to our eyes they may overlap each other in terms of class memberships. Namely, visualization of vectors in much lower dimensions may show overlap among these vectors.
- However, when viewed in their ambient space \(\mathbb{R}^{p}\), they may not overlap at all (since neighborhoods of a vector are different in spaces of different dimensions).
- Think about an overpass on a highway, which “intersects” the highway when both are viewed from above (equivalent to visualizing them in 2D)
LDA: p > 2n
LDA with 50 feature variables, 3 classes, and sample size 20:
> library(MASS)
> lda.fit = lda(Class~.,data=tmp)
Warning in lda.default(x, grouping, ...): variables are
collinear
> lda.pred = predict(lda.fit,tmp)
> TrueClassLabel=tmp$Class
> LDAEstimtedClassLabel=lda.pred$class
> table(LDAEstimtedClassLabel,TrueClassLabel)
TrueClassLabel
LDAEstimtedClassLabel BREAST COLON LEUKEMIA
BREAST 6 0 0
COLON 1 7 0
LEUKEMIA 0 0 6Classwise error rates: “BREAST”, “LEUKEMIA” and “COLON”
[1] 0.1428571 0.0000000 0.0000000- Standard estimates of covariance matrix are singular!!
QDA: p > 2n
The component covariance matrices may not be equal since gene profiles for different cancer types may have different dependence structures. So, let us apply QDA:
> library(MASS)
> qda.fit = qda(Class~.,data=tmp)
> # Error in qda.default(x, grouping, ...) :
> # some group is too small for 'qda'- Not enough observations in each group to sensibly estimate the corresponding component parameters
- Standard estimates of component covariance matrices are singular!!
Singular DA: p > n
- LDA: pooled covariance matrix estimate \[ \hat{\Sigma}=\frac{1}{n-K}\sum_{k=1}^{K}\sum_{\left\{ i:y_{i}=k\right\} }\left( x_{i}-\hat{\mu}_{k}\right) \left( x_{i}-\hat{\mu}_{k}\right) ^{T} \] as an estimate of \(\Sigma\)
- QDA: sample covariance matrix \[ \hat{\Sigma}_k = \frac{1}{n_k-1}\sum_{\left\{ i:y_{i}=k\right\} }\left( x_{i}-\hat{\mu}_{k}\right) \left( x_{i}-\hat{\mu}_{k}\right) ^{T} \] as an estimate of \(\Sigma_k\)
Caution: when \(p >n\), \(\hat{\Sigma}\) and all \(\hat{\Sigma}_k\) are all singular, not being valid covariance matrices or forcing some feature variables to be linearly dependent!
Regularized DA
When \(p >n\), we can use the regularized estimate proposed by “Friedman, J.H. (1989): Regularized Discriminant Analysis” and implemented by R library
klaR.The regularized estimate of \(\Sigma_k\) is \[ \hat{\Sigma}_{k,\lambda,\gamma}= (1-\gamma) \hat{\Sigma}_{k,\lambda} + \gamma p^{-1} \text{tr}(\hat{\Sigma}_{k,\lambda}) \mathbf{I}, \gamma \in [0,1] \] with \[\hat{\Sigma}_{k,\lambda}=(1-\lambda) \hat{\Sigma}_k + \lambda \hat{\Sigma}, \lambda \in [0,1]\] where \(\hat{\Sigma}\) is a pooled covariance matrix, \(\hat{\Sigma}_k\) is a suitable estimate of \(\Sigma_k\), and \(\text{tr}(A)\) is the trace of the square matrix \(A\), i.e., the sum of the diagonal entries of \(A\)
Regularized DA via klaR
We need R library klaR, and command rda{klaR} for which the basic syntax is
rda(formula, data, gamma = 0.05, lambda = 0.2,
crossval = FALSE, ...)formula: the same as inldaorqdagamma,lambda: same as \(\gamma\) and \(\lambda\) in \(\hat{\Sigma}_{k,\lambda,\gamma}\). If unspecified, either or both are determined by minimizing the estimated error ratecrossval: logical. IfTRUE, then the error rate is estimated by Cross-Validation; otherwise, by drawing several training- and test-samples.
Regularized DA via klaR
rda{klaR} returns a list of class rda containing the following components:
regularization: vector containing the two regularization parameters (gamma, lambda).classesandprior: the names of the classes, and the prior probabilities for the classes.error.rate: apparent error rate (if computation was not suppressed), and, if any optimization took place, the final (cross-validated or bootstrapped) error rate estimate as well.
Regularized DA via klaR
rda{klaR} returns a list of class rda containing the following components:
means,covariancesandcovpooled: group means, array of group covariances, and pooled covariance.converged: (Logical) indicator of convergence (only for Nelder-Mead).iter: number of iterations actually performed (only for Nelder-Mead).
Regularized DA via klaR
RDA on the subset data (\(p=50\) and \(n=20\)):
> library(klaR)
> rda.fit = rda(Class~.,data=tmp, gamma = 0.05, lambda = 0.2)
> rda.pred = predict(rda.fit, tmp) # classify obs in `tmp`
> TrueClassLabel=tmp$Class
> rDAEstimtedClassLabel=rda.pred$class
> table(rDAEstimtedClassLabel,TrueClassLabel)
TrueClassLabel
rDAEstimtedClassLabel BREAST COLON LEUKEMIA
BREAST 7 0 0
COLON 0 7 0
LEUKEMIA 0 0 6predictacts on an object of classrdasimilarly asMASS::predicton an object of classldaorqda- good results; not determining
gammaorlambdavia cross-validation due ton=20
Extract est. covariance matrices
> # extract estimated covariance matrices
> hatSigma1 = rda.fit$covariances[,,1]
> hatSigma2 = rda.fit$covariances[,,2]
> hatSigma3 = rda.fit$covariances[,,3]rda(...)$covariances: a \(p \times p \times K\) array. To obtain the estimated covariance matrix, \(\hat{\Sigma}_{k,\lambda,\gamma}\), of observations in Class \(k\), userda(...)$covariances[,,k]. Note that class (and label) ordering is determined byrdausing rules of R.
Melt est. covariance matrices
> # convert estimated covariance matrices for plotting
> library(reshape2)
> melted_hatSigma1 = melt(hatSigma1)
> melted_hatSigma2 = melt(hatSigma2)
> melted_hatSigma3 = melt(hatSigma3)
> EstSigmaAll=rbind(melted_hatSigma1,melted_hatSigma2,
+ melted_hatSigma3)
> EstSigmaAll$Cancer=rep(c("BREAST","COLON","LEUKEMIA"),
+ each=nrow(melted_hatSigma1))
> EstSigmaAll$Cancer=factor(EstSigmaAll$Cancer)melt(data,value.name ="value",...): convertdatainto amolten data.frame, and by default store entries ofdataas"value"in themolten data.frame, as if values indataare values of a function that are obtained on a 2D grid on two variables
Plot est. covariance matrices
> library(ggplot2)
> ggplot(data=EstSigmaAll, aes(x=Var1, y=Var2, fill=value)) +
+ geom_tile()+scale_fill_gradient2(low="blue", high="red")+
+ facet_grid(~Cancer)+xlab("")+ylab("")+
+ ggtitle("Estimated covariance matrices")+
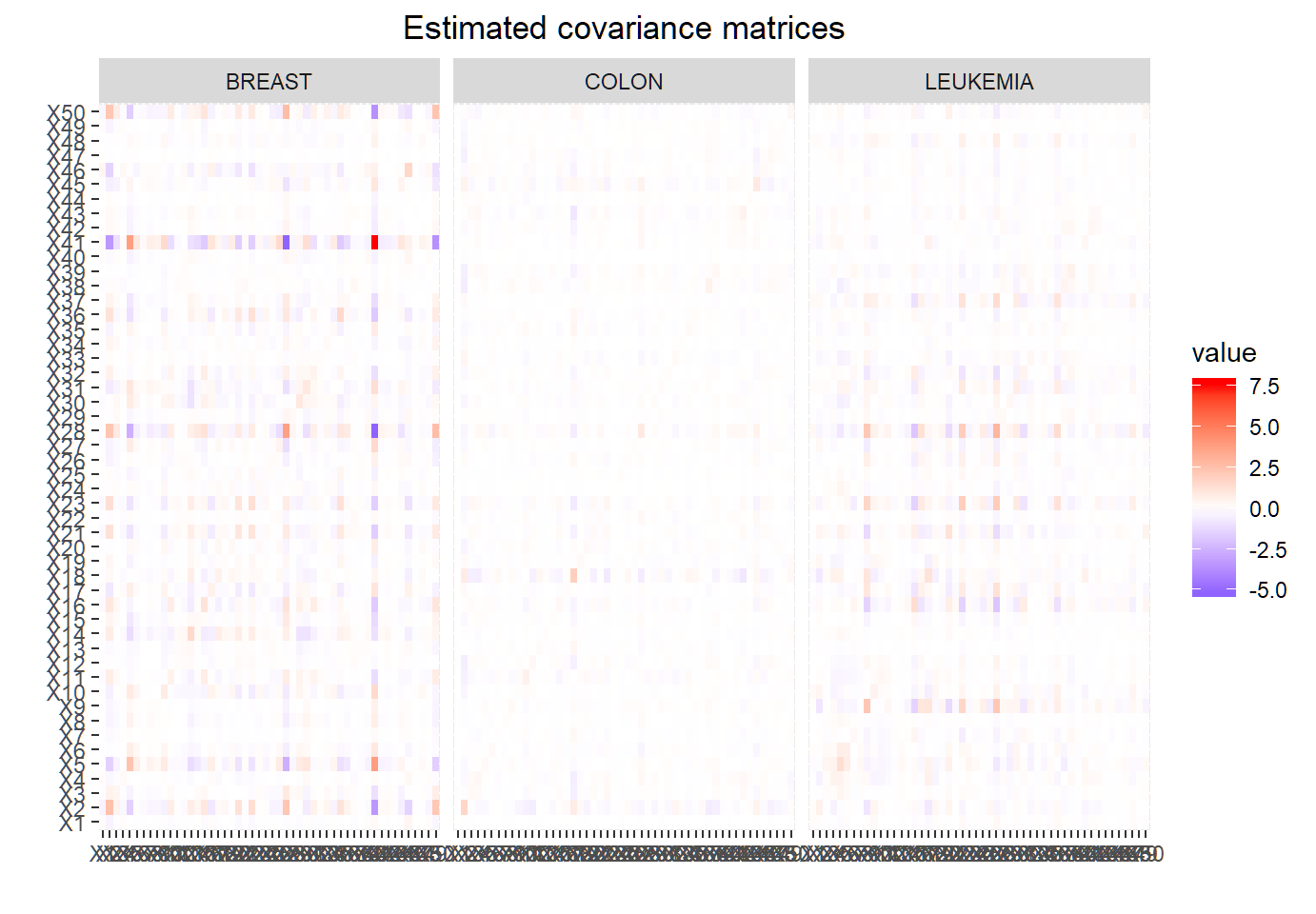
+ theme(plot.title = element_text(hjust = 0.5))ggplot(...,aes(x=, y=, fill=value))should be used withxlab("")+ylab("")since no x-axis or y-axis label is needed- feature variables will be shown on both axes since pairwise covariances are computed for them
- resulting plot is similar to a heatmap
Visualization of est. cov. matrx.
Much less dependence among genes for “COLON” cancer than for “BREAST” or “LEUKEMIA” cancer
Discriminant analysis: Example 4
Stock market data
Data set Smarket{ISLR} contains observations for S&P 500 stock index over 1,250 days, from the beginning of 2001 until the end of 2005, such as
- percentage returns for each of the five previous trading days, as
Lag1throughLag5 Direction(whether market wasUporDownon this date)
> library(ISLR); dim(Smarket)
[1] 1250 9
> Smarket[1,]
Year Lag1 Lag2 Lag3 Lag4 Lag5 Volume Today Direction
1 2001 0.381 -0.192 -2.624 -1.055 5.01 1.1913 0.959 Up
> levels(Smarket$Direction)
[1] "Down" "Up" Target: predict market Direction using Lag1 and Lag2
LDA: training
Model: 2 features \(X_1=\)Lag1 and \(X_2=\)Lag2; 2 component bivariate Gaussian densities \(f_1 \sim \text{Gaussian}(\mu_1,\Sigma_1)\) and \(f_2 \sim \text{Gaussian}(\mu_2,\Sigma_2)\); 2 mixing proportions (as priors) \(\pi_1\) and \(\pi_2\); 2 classes Down and Up
> train=(Smarket$Year<2005); Smarket.2005=Smarket[!train,]
> Direction.2005=Smarket$Direction[!train]
> library(MASS)
> lda.fit=lda(Direction~Lag1+Lag2,data=Smarket,subset=train)- observations from 2001 through 2004 as training set
- note the use of
subset=traininlda - LDA assumes \(\Sigma_1=\Sigma_2\)
LDA: estimated model
Estimated prior (i.e., mixing) probabilities and mean vectors:
Prior probabilities of groups:
Down Up
0.491984 0.508016
Group means:
Lag1 Lag2
Down 0.04279022 0.03389409
Up -0.03954635 -0.03132544- Class 1 is
Direction=Down; Class 2 isDirection=Up - \(\hat{\pi}_1=0.492\), \(\hat{\pi}_2=0.508\), \(\hat{\mu}_1=(0.0428,0.0339)^T\), \(\hat{\mu}_2=(-0.0395,-0.0313)^T\)
LDA: estimated model
Recall the estimated linear discriminant function as \[\hat{\delta}_{k}(x)=x^T \underbrace{\hat{\Sigma}^{-1}\hat{\mu}_k}_{\hat{c}_k} -\frac{1}{2} \hat{\mu}_k^T \underbrace{\Sigma^{-1}\hat{\mu}_k}_{\hat{c}_k} + \log \hat{\pi}_k, k \in \mathcal{G}\]
Coefficients of linear discriminants:
LD1
Lag1 -0.6420190
Lag2 -0.5135293- \(\hat{c}_2=(-0.642,-0.514)^T\), since Class 2 is
Direction=UpandUp\(\mapsto 1\) (and Class 1 asDirection=DownandDown\(\mapsto 0\))
Linear discriminants
Define estimated linear discriminant as \(\hat{d}_{k}(x)=x^T \hat{c}_k,k \in \mathcal{G}\)
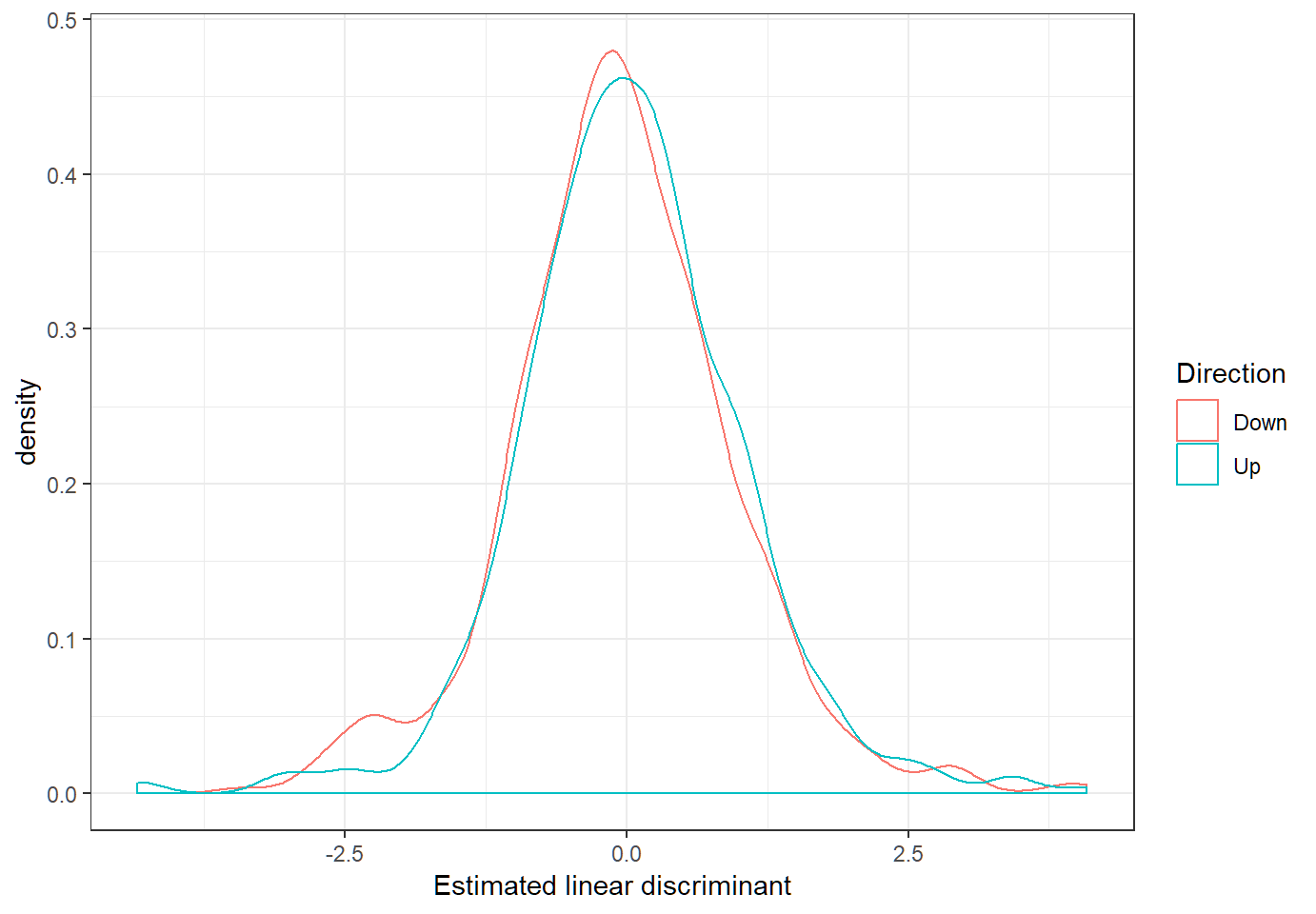
LDA: predicting
Classification on test set Smarket.2005:
> dim(Smarket.2005)
[1] 252 9
> lda.pred=predict(lda.fit, Smarket.2005)
> names(lda.pred)
[1] "class" "posterior" "x"
> lda.class=lda.pred$class
> table(lda.class,Direction.2005)
Direction.2005
lda.class Down Up
Down 35 35
Up 76 106
> mean(lda.class==Direction.2005)
[1] 0.5595238lda.pred$x: vector of estimated linear discriminant for each observation- classwise error rates:
DownandUprespectively
[1] 0.6846847 0.2482270QDA: estimated model
> qda.fit=qda(Direction~Lag1+Lag2,data=Smarket,subset=train)
> qda.fit
Call:
qda(Direction ~ Lag1 + Lag2, data = Smarket, subset = train)
Prior probabilities of groups:
Down Up
0.491984 0.508016
Group means:
Lag1 Lag2
Down 0.04279022 0.03389409
Up -0.03954635 -0.03132544qdadoes not produce “coefficients of linear discriminants” since it uses a quadratic decision boundary- estimated prior (i.e., mixing) probabilities and mean vectors are the same as those obtained by LDA
QDA: predicting
Classification on test set Smarket.2005:
> qda.class=predict(qda.fit,Smarket.2005)$class
> table(qda.class,Direction.2005)
Direction.2005
qda.class Down Up
Down 30 20
Up 81 121
> mean(qda.class==Direction.2005)
[1] 0.5992063- around 60% (average) accuracy of QDA, in comparison with around 56% (average) accuracy of LDA
- QDA: impressive performance for these stock market data
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19041)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] highr_0.7 stringr_1.3.1 xfun_0.4
[13] digest_0.6.18 evaluate_0.12