Stat 437 Lecture Notes 6b
Xiongzhi Chen
Washington State University
Support vector machines: software implementation
R software and commands
R libraries and commands needed:
- R library
e1071, commandsvm{e1071}to construct a support vector machine (SVM), and commandtune{e1071}to determine cost parameter \(C\) (and kernel parameters) by cross-validation - R library
LiblineaRfor very large linear problems (not discussed here)
Command svm{e1071}
svm{e1071} is used to train a support vector machine. Its basic syntax is
svm(formula, data=NULL, subset, scale=TRUE, kernel="radial",
degree=3, gamma=if (is.vector(x)) 1 else 1/ncol(x),
cost=1, na.action=na.omit, ...)formula: a symbolic description of the model to be fit.data: an optional data frame containing the variables in the model. By default the variables are taken from the environment whichsvmis called from.subset: an index vector specifying the cases to be used in the training sample. (NOTE: If given, this argument must be named.)
Command svm{e1071}
Basic syntax:
svm(formula, data=NULL, subset, scale=TRUE, kernel="radial",
degree=3, gamma=if (is.vector(x)) 1 else 1/ncol(x),
cost=1, na.action=na.omit, ...)scale: a logical vector indicating the variables to be scaled. By default, data are scaled internally (bothxandyvariables) to zero mean and unit variance. The center and scale values are returned and used for later predictions.kernel: the kernel used in training and predicting, which can belinear(i.e., the Euclidean inner product),polynomialwithdegree, orradial(i.e., the radial kernel).
Command svm{e1071}
Basic syntax:
svm(formula, data=NULL, subset, scale=TRUE, kernel="radial",
degree=3, gamma=if (is.vector(x)) 1 else 1/ncol(x),
cost=1, na.action=na.omit, ...)degree: degree ofpolynomialkernel; default is 3.gamma: parameter needed forradialkernel; default is \(1/p\) (where \(p\) is the number of features).cost: cost of constraints on violation, i.e., the cost \(C\) in the Lagrange multiplier function; default is 1.
Command svm{e1071}
svm{e1071} returns an object of class svm containing the fitted model, including:
SV: support vectors (possibly scaled).index: indices of support vectors in the data matrix. Note that thisindexrefers to the preprocessed data (after the possible effect ofna.omitandsubset).coefs: coefficients (i.e., entries of the direction vector of optimal hyperplane) times the training labels.rho: the negative intercept (of optimal hyperplane).
Command tune{e1071}
tune{e1071} tunes hyperparameters of statistical methods using a grid search over supplied parameter ranges. Its basic syntax is:
tune(method, data = list(), ranges = NULL, ...)method: either the function to be tuned, or a character string naming such a function.data: data, if a formula interface is used; ignored, if predictor matrix and response are supplied directly.ranges: a named list of parameter vectors spanning the sampling space. The vectors will usually be created byseq.
Command tune{e1071}
Basic syntax:
tune(method, data = list(), ranges = NULL, ...)...: additional parameters and some parameters ofmethod
tune{e1071} returns an object of class tune, including the components:
best.parameters: a1 x kdata frame withkas number of parameters.best.performance: best achieved performance.
Support vector classifier (SVC): Example 1a
Nonseparable data
Generate nonseparable data:
- \(n=20\) observations on \(p=2\) features \(X_1,X_2\)
- Class \(-1\): \(x_i \sim \text{Gaussian}(\mu_1=(0,0)^T,\mathbf{I}),i=1,\ldots,10\)
- Class \(1\): \(x_i \sim \text{Gaussian}(\mu_2=(1,1)^T,\mathbf{I}),i=11,\ldots,20\)
> set.seed(1)
> x=matrix(rnorm(20*2), ncol=2)
> y=c(rep(-1,10), rep(1,10))
> x[y==1,]=x[y==1,] + 1
> dat=data.frame(x=x, y=as.factor(y))Scatter plot
No separating hyperplane:
> par(mar = c(4.9, 4.5, .0, .5),oma=c(1.3,1.3,0.3,1.3))
> plot(x,col=(3-y),xlab=expression(X[1]),ylab=expression(X[2]))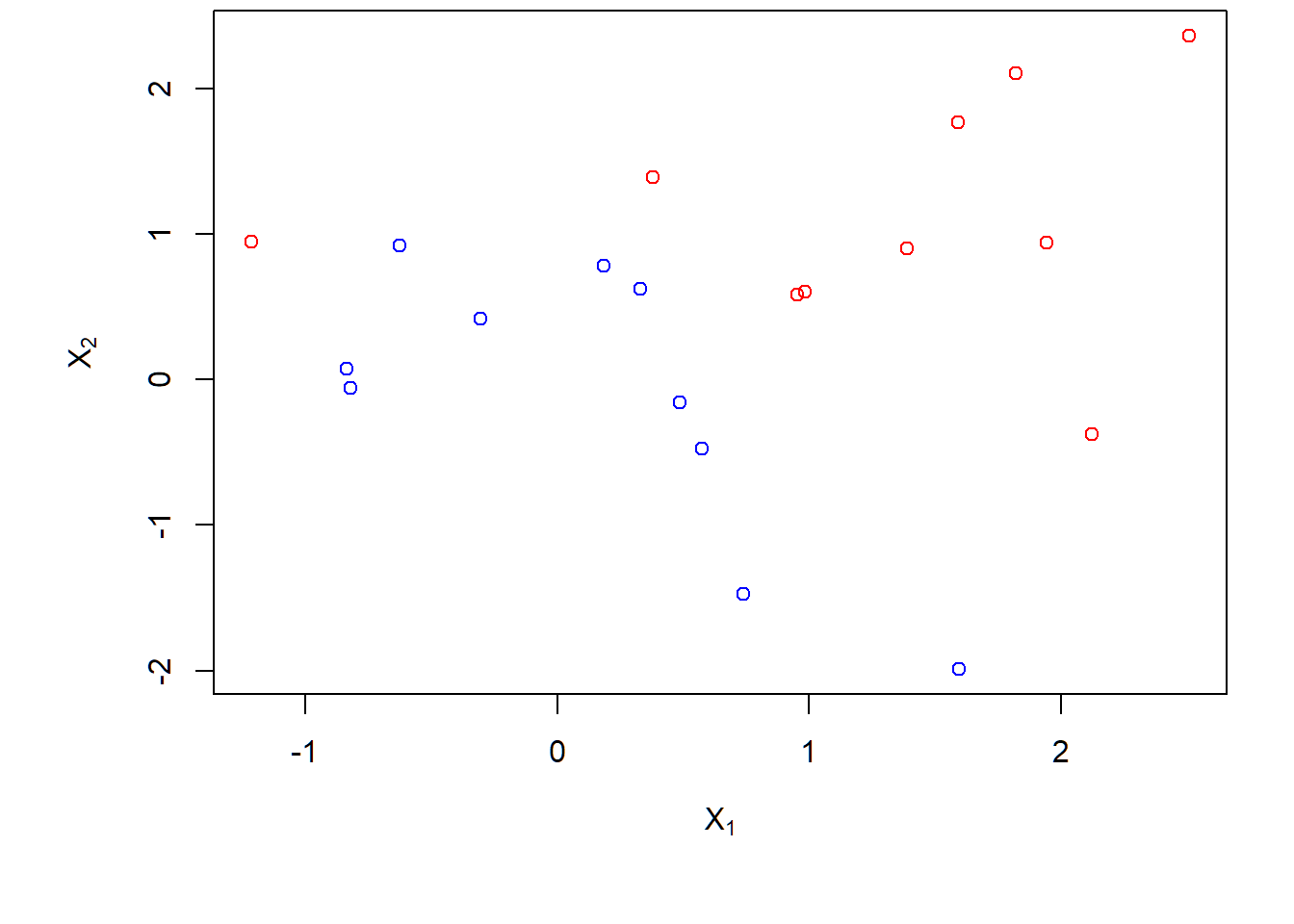
SVC: \(C=10\)
SVC with cost \(C=10\):
> library(e1071)
> svmfit=svm(y~.,data=dat,kernel="linear",cost=10,scale=FALSE)
> svmfit$index # indices of support vectors
> summary(svmfit)
> plot(svmfit, dat) # X_2 on x-axisscaleasTRUEorFALSEdependent on scales of measurements on feature variablessummary(svm.ojb): provide a summary of model and resultsplot(svm.ojb,data): create visualization of classification results by plotting 2nd feature on x-axis
SVC: \(C=10\)
Summary of model fit:
[1] 1 2 5 7 14 16 17
Call:
svm(formula = y ~ ., data = dat, kernel = "linear",
cost = 10, scale = FALSE)
Parameters:
SVM-Type: C-classification
SVM-Kernel: linear
cost: 10
gamma: 0.5
Number of Support Vectors: 7
( 4 3 )
Number of Classes: 2
Levels:
-1 1- 4 support vectors in one class and 3 in the other
SVC: \(C=10\)
Support vectors as crosses; others observations as circles
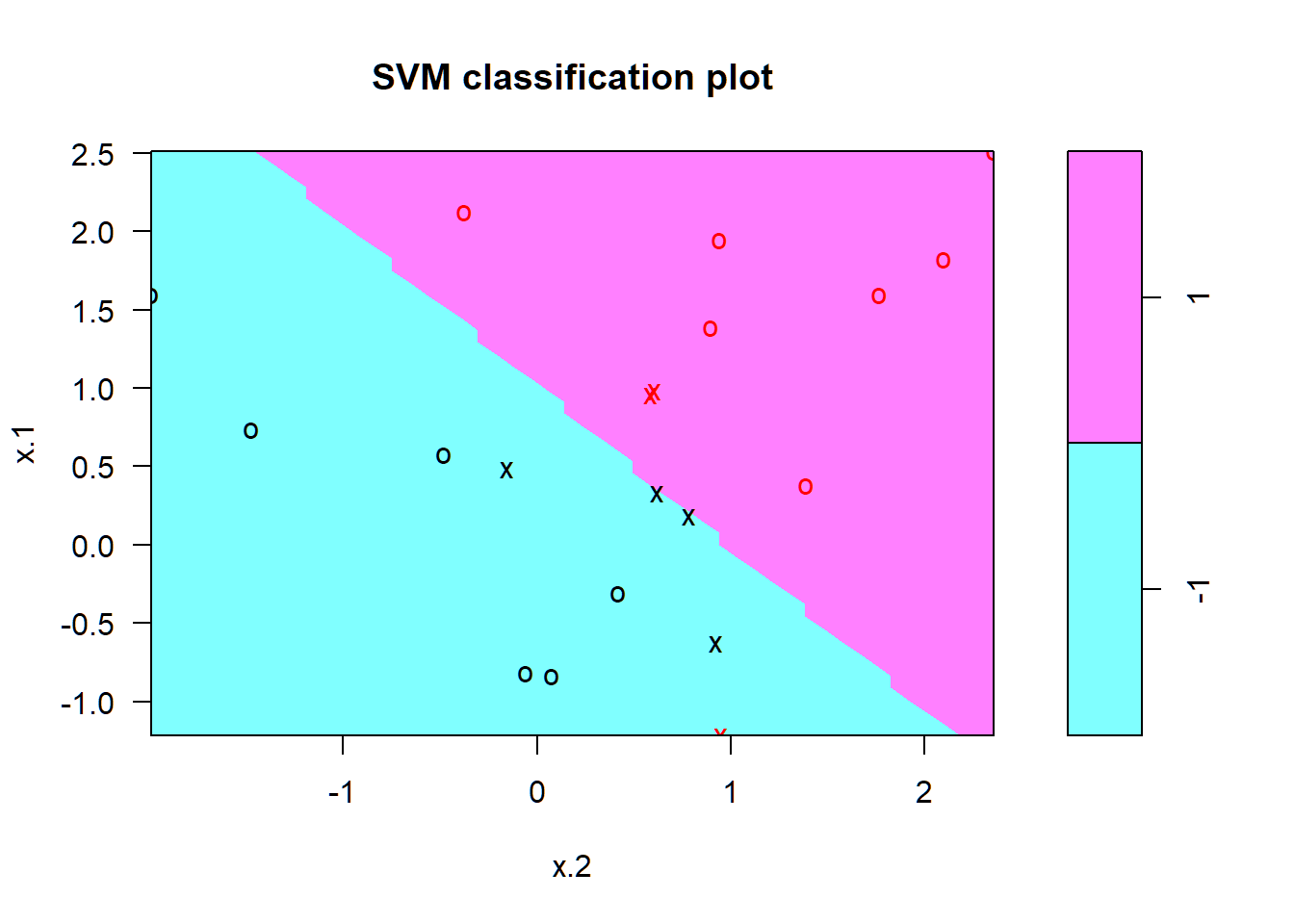
SVC: \(C=0.1\)
> svmfit=svm(y~.,data=dat,kernel="linear",cost=0.1,scale=FALSE)
> svmfit$index
[1] 1 2 3 4 5 7 9 10 12 13 14 15 16 17 18 20
> par(mar = c(4.9, 4.5, 1.8, .5),oma=c(1.3,1.3,0.3,1.3)); plot(svmfit, dat)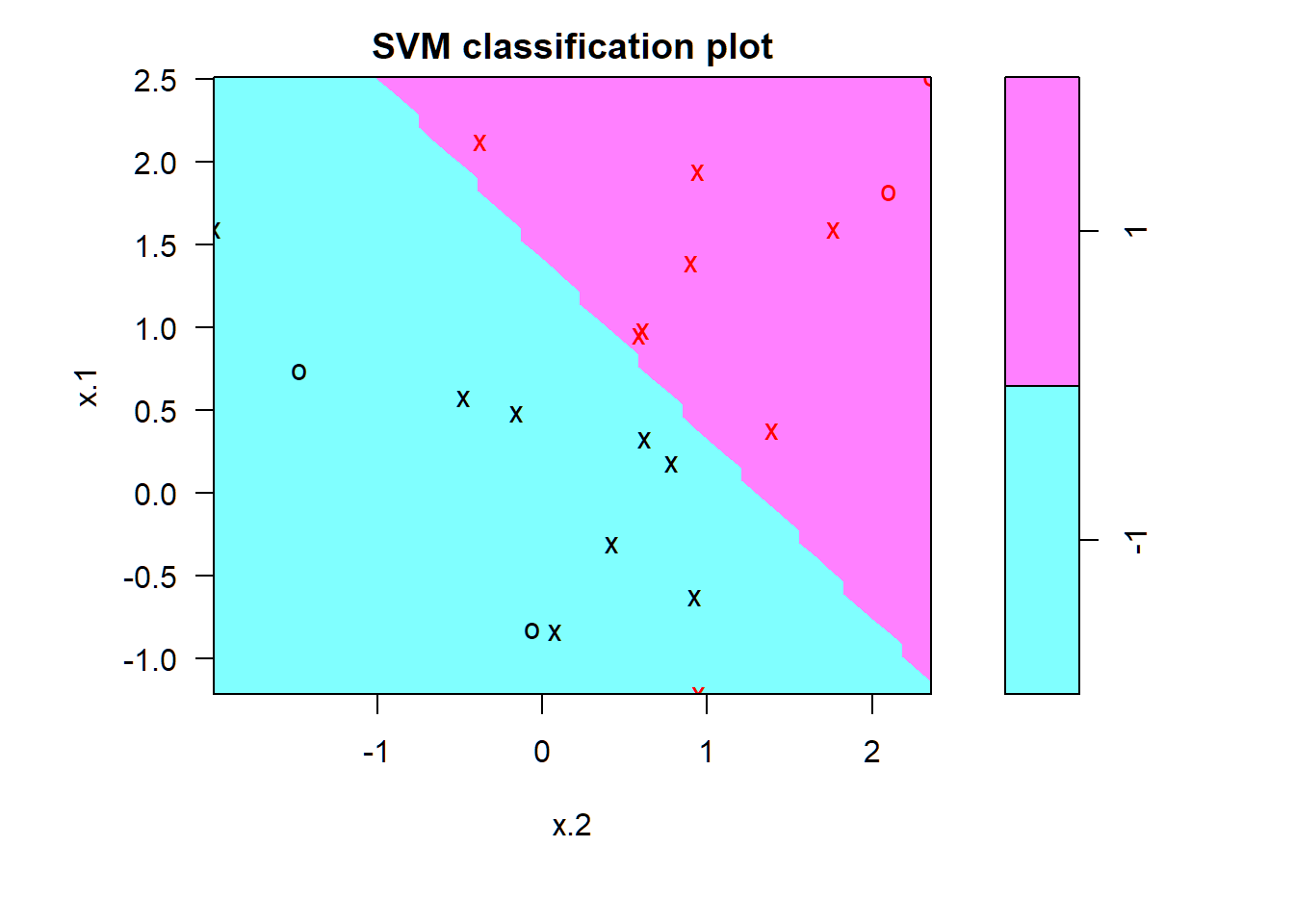
Cross-validation on \(C\)
> set.seed(1)
> tune.out=tune(svm,y~.,data=dat,kernel="linear",
+ ranges=list(cost=c(0.001, 0.01, 0.1, 1,5,10,100)))
> summary(tune.out)
Parameter tuning of 'svm':
- sampling method: 10-fold cross validation
- best parameters:
cost
0.1
- best performance: 0.1
- Detailed performance results:
cost error dispersion
1 1e-03 0.70 0.4216370
2 1e-02 0.70 0.4216370
3 1e-01 0.10 0.2108185
4 1e+00 0.15 0.2415229
5 5e+00 0.15 0.2415229
6 1e+01 0.15 0.2415229
7 1e+02 0.15 0.2415229set.seedneeded since cross-validation randomly divides data into splits of approximately equal size
Obtain best model
Obtain best model given by cross-validation: \(C=0.1\)
> bestmod=tune.out$best.model
> summary(bestmod)
Call:
best.tune(method = svm, train.x = y ~ ., data = dat,
ranges = list(cost = c(0.001, 0.01, 0.1, 1, 5,
10, 100)), kernel = "linear")
Parameters:
SVM-Type: C-classification
SVM-Kernel: linear
cost: 0.1
gamma: 0.5
Number of Support Vectors: 16
( 8 8 )
Number of Classes: 2
Levels:
-1 1tune.obj$best.model: best model given by cross-validation
Best model applied to test set
Generate test set:
- \(n=20\) observations on \(p=2\) features \(X_1,X_2\)
- Class \(-1\): \(x_i \sim \text{Gaussian}(\mu_1=(0,0)^T,\mathbf{I})\)
- Class \(1\): \(x_j \sim \text{Gaussian}(\mu_2=(1,1)^T,\mathbf{I})\)
> set.seed(1)
> xtest=matrix(rnorm(20*2), ncol=2)
> # create class labels
> ytest=sample(c(-1,1), 20, rep=TRUE)
> xtest[ytest==1,]=xtest[ytest==1,] + 1
> testdat=data.frame(x=xtest, y=as.factor(ytest))
> ypred=predict(bestmod,testdat)
> table(predict=ypred, truth=testdat$y)
truth
predict -1 1
-1 10 1
1 1 8- note the syntax
predict(svm.ojb,data)
SVC: \(C=0.01\)
SVC with \(C=0.01\) on training data:
> svmfit=svm(y~.,data=dat,kernel="linear",cost=.01,scale=FALSE)
> ypred1=predict(svmfit,testdat)
> table(predict=ypred1, truth=testdat$y)
truth
predict -1 1
-1 10 4
1 1 5- Caution: not able to reproduce Text results for this
- \(C=0.01\) not as good as \(C=0.1\)
Support vector classifier (SVC): Example 1b
Barely separable data
Observations barely separable by a hyperplane:
> x[y==1,]=x[y==1,]+0.5; dat=data.frame(x=x,y=as.factor(y))
> par(mar=c(6.2,4.5,.0,.5),oma=c(1.3,1.3,0.3,1.3))
> plot(x, col=(y+5)/2, pch=19,
+ xlab=expression(X[1]),ylab=expression(X[2]))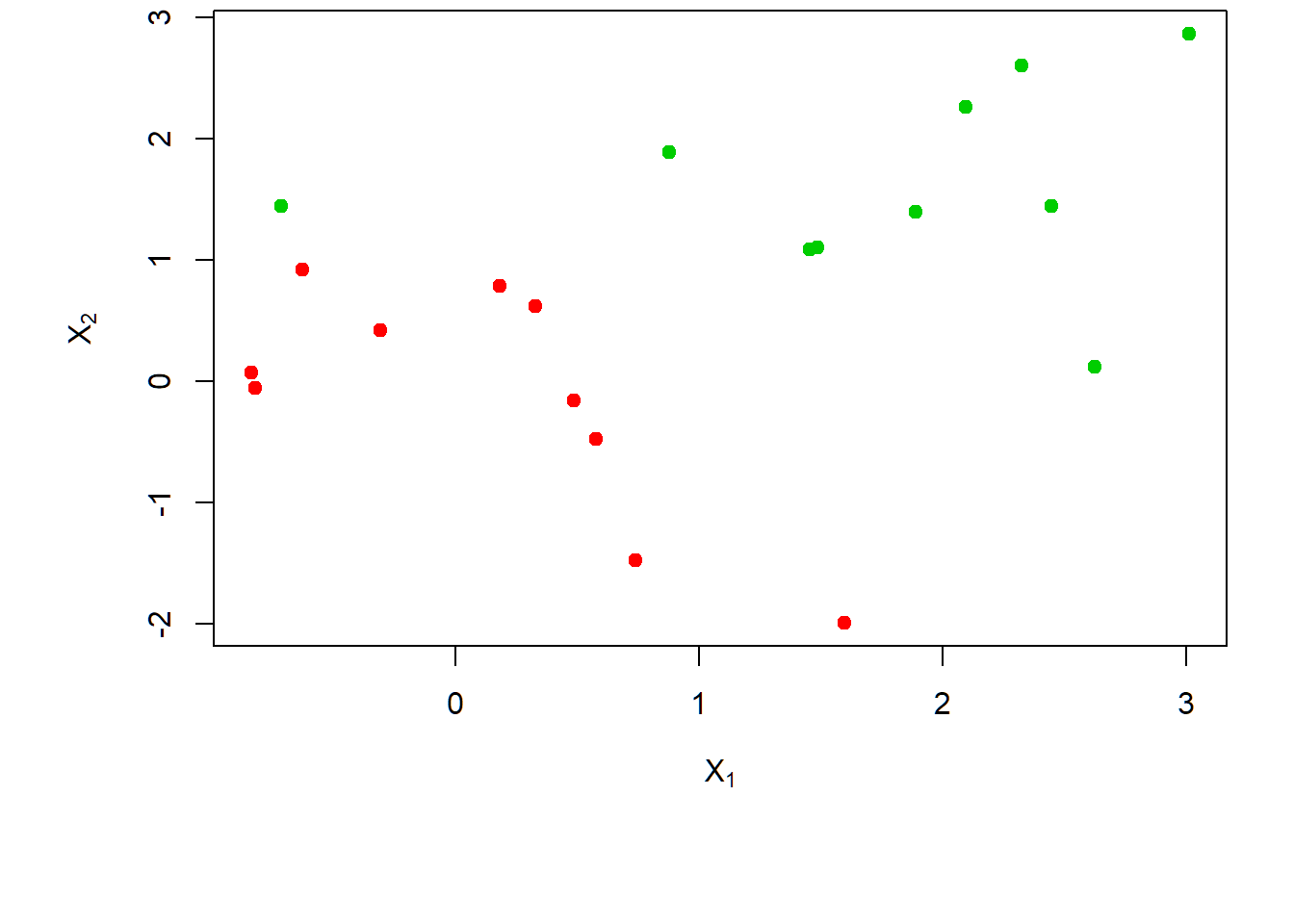
SVC: \(C=10^{5}\)
SVC with cost \(C=10^{5}\):
> svmfit=svm(y~., data=dat, kernel="linear", cost=1e5)
> summary(svmfit)
Call:
svm(formula = y ~ ., data = dat, kernel = "linear",
cost = 1e+05)
Parameters:
SVM-Type: C-classification
SVM-Kernel: linear
cost: 1e+05
gamma: 0.5
Number of Support Vectors: 3
( 1 2 )
Number of Classes: 2
Levels:
-1 1- \(C\) very large so that no observations are misclassified
SVC: \(C=10^{5}\)
Zero training error; fitted model may overfit training set and not perform well on test set
> par(mar=c(7,4.5,1.2,.5),oma=c(1.3,1.3,0.3,1.3)); plot(svmfit, dat)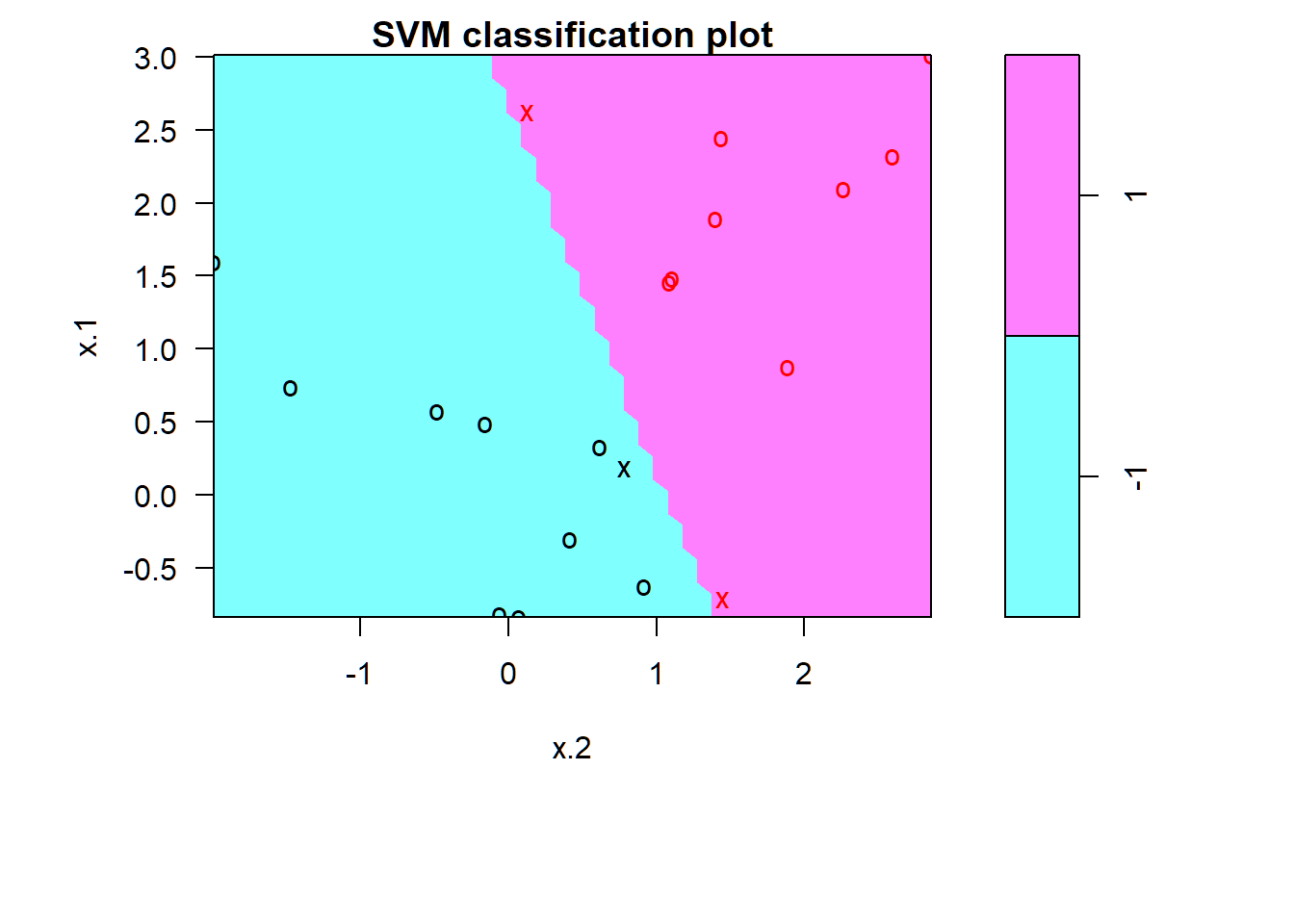
SVC: \(C=1\)
SVC with cost \(C=1\):
> svmfit=svm(y~., data=dat, kernel="linear", cost=1)
> summary(svmfit)
Call:
svm(formula = y ~ ., data = dat, kernel = "linear",
cost = 1)
Parameters:
SVM-Type: C-classification
SVM-Kernel: linear
cost: 1
gamma: 0.5
Number of Support Vectors: 7
( 4 3 )
Number of Classes: 2
Levels:
-1 1SVC: \(C=1\)
1 observation misclassified; fitted model may perform well on test set
> par(mar=c(7,4.5,1.2,.5),oma=c(1.3,1.3,0.3,1.3)); plot(svmfit,dat)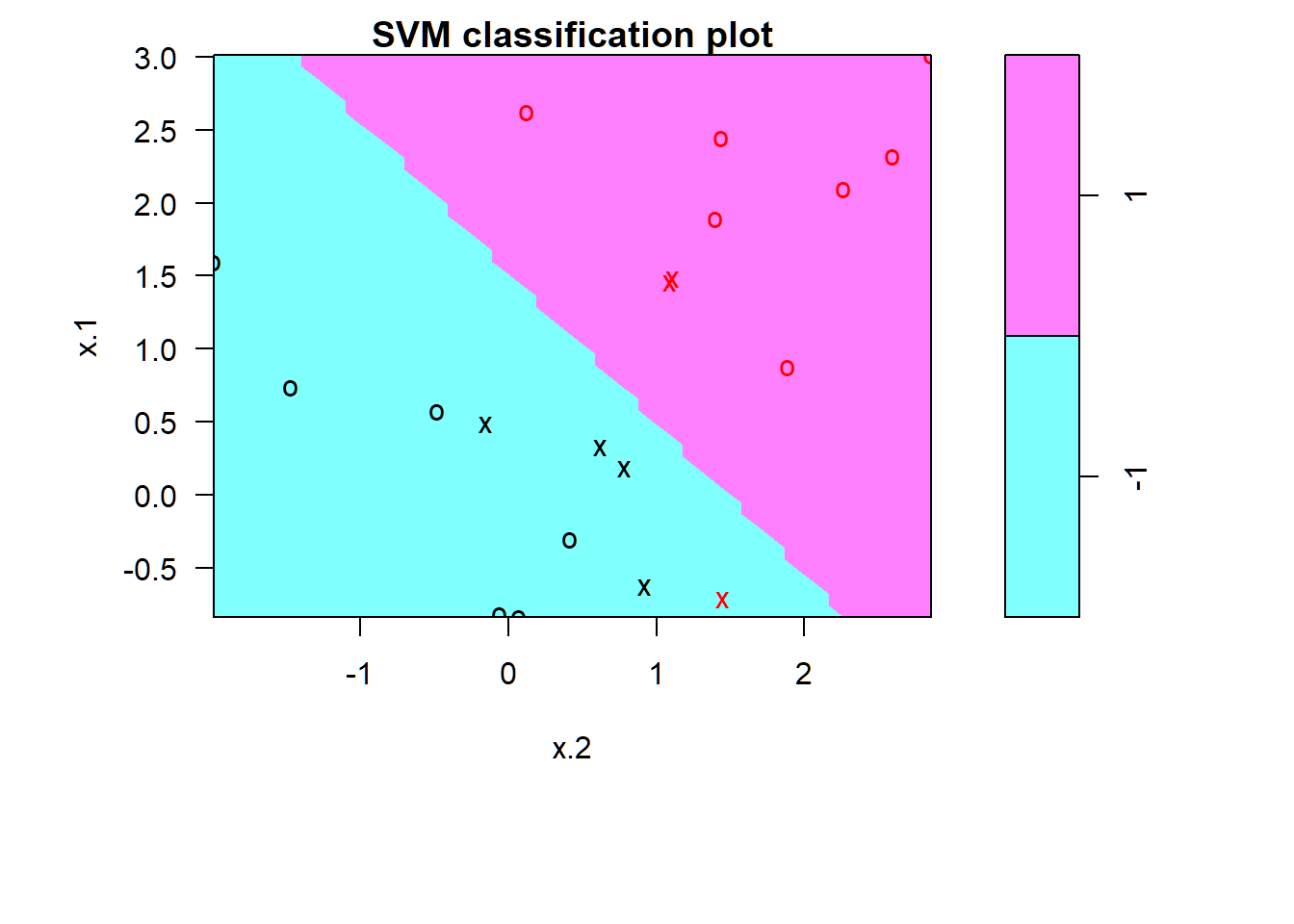
Support vector machines (SVMs): Example 1
Generate data
Data with a nonlinear class boundary:
- \(n=200\) observations on \(p=2\) features \(X_1,X_2\)
- Class \(1\): \(150\) observations; \[x_i \sim \text{Gaussian}(\mu_1=(2,2)^T,\mathbf{I}),i=1,\ldots,100\\ x_i \sim \text{Gaussian}(\mu_2=(-2,-2)^T,\mathbf{I}),i=101,\ldots,150 \]
- Class \(2\): \(50\) observations; \(x_i \sim \text{Gaussian}(\mu_3=(0,0)^T,\mathbf{I}),i=151,\ldots,200\)
> set.seed(1)
> x=matrix(rnorm(200*2), ncol=2)
> x[1:100,]=x[1:100,]+2
> x[101:150,]=x[101:150,]-2
> y=c(rep(1,150),rep(2,50))
> dat=data.frame(x=x,y=as.factor(y))Scatter plot
Data with a nonlinear class boundary:
> par(mar=c(7,4.5,1.2,.5),oma=c(1.3,1.3,0.3,1.3))
> plot(x, col=y,xlab=expression(X[1]),ylab=expression(X[2]))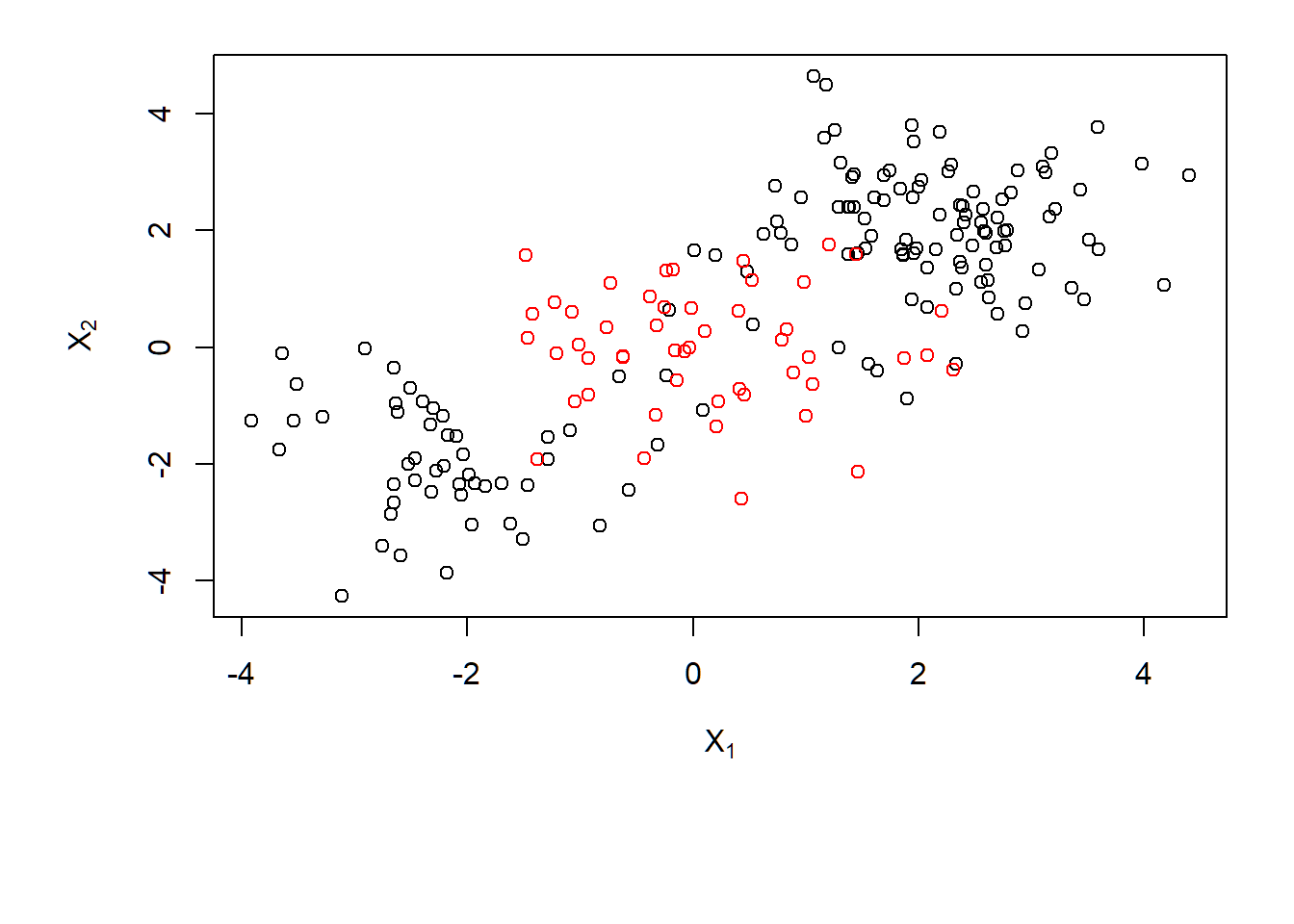
SVM: radial kernel; \(C=1\)
SVM with radial kernel and \(\gamma=1\) and cost \(C=1\) on training set of \(100\) observations:
> train=sample(200,100) # training obs index
> svmfit=svm(y~., data=dat[train,], kernel="radial",
+ gamma=1, cost=1)
> summary(svmfit)
Call:
svm(formula = y ~ ., data = dat[train, ], kernel = "radial",
gamma = 1, cost = 1)
Parameters:
SVM-Type: C-classification
SVM-Kernel: radial
cost: 1
gamma: 1
Number of Support Vectors: 37
( 17 20 )
Number of Classes: 2
Levels:
1 2SVM: radial kernel; \(C=1\)
Support vectors as crosses; other observations as circles; fair number of training errors:
> par(mar=c(7,4.5,1.4,.5),oma=c(1.3,1.3,0.3,1.3)); plot(svmfit, dat[train,])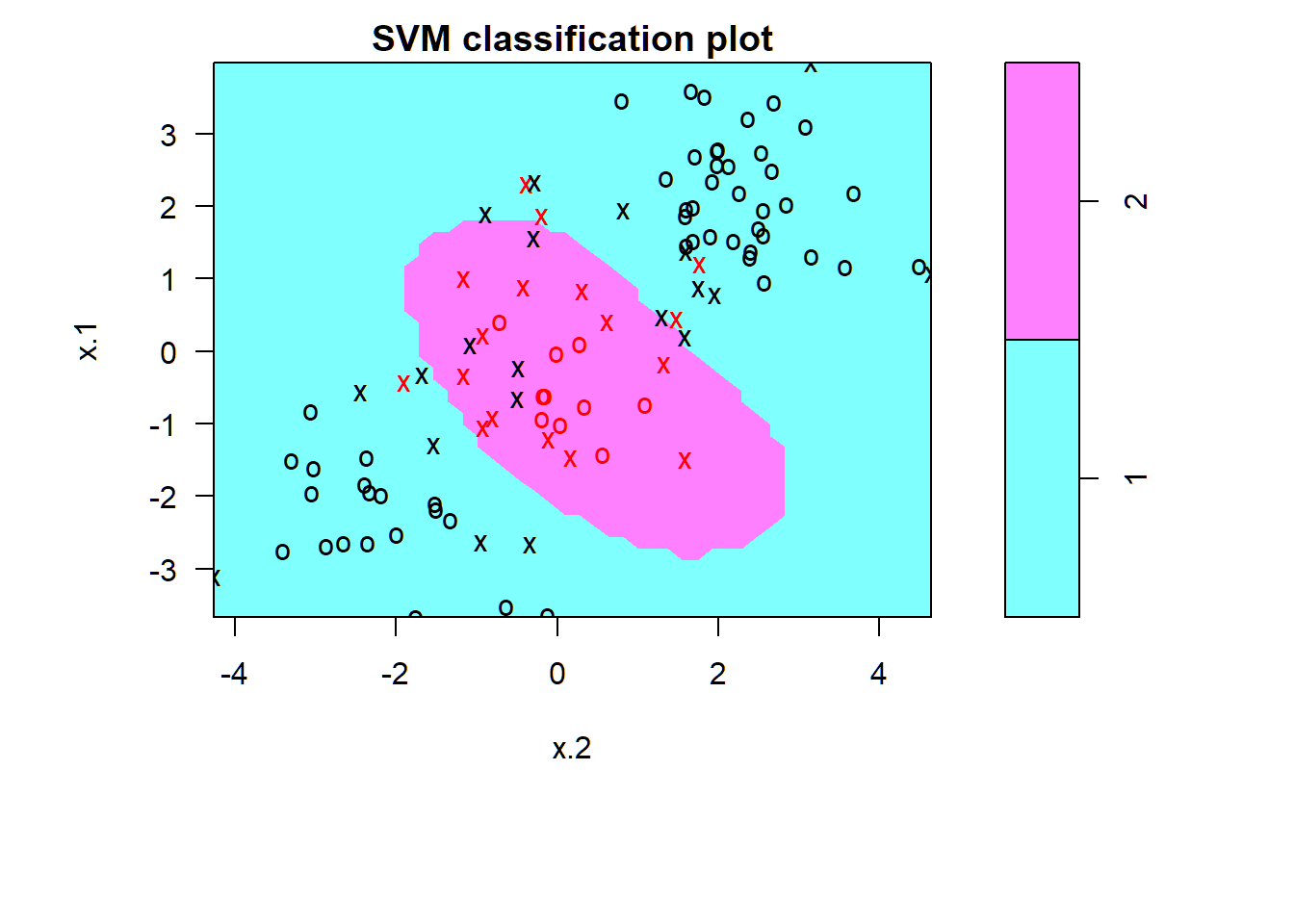
SVM: radial kernel; \(C=10^5\)
Less training error; risk of overfitting training data
> svmfit=svm(y~., data=dat[train,], kernel="radial",
+ gamma=1,cost=1e5)
> par(mar=c(7,4.5,1.4,.5),oma=c(1.3,1.3,0.3,1.3)); plot(svmfit,dat[train,])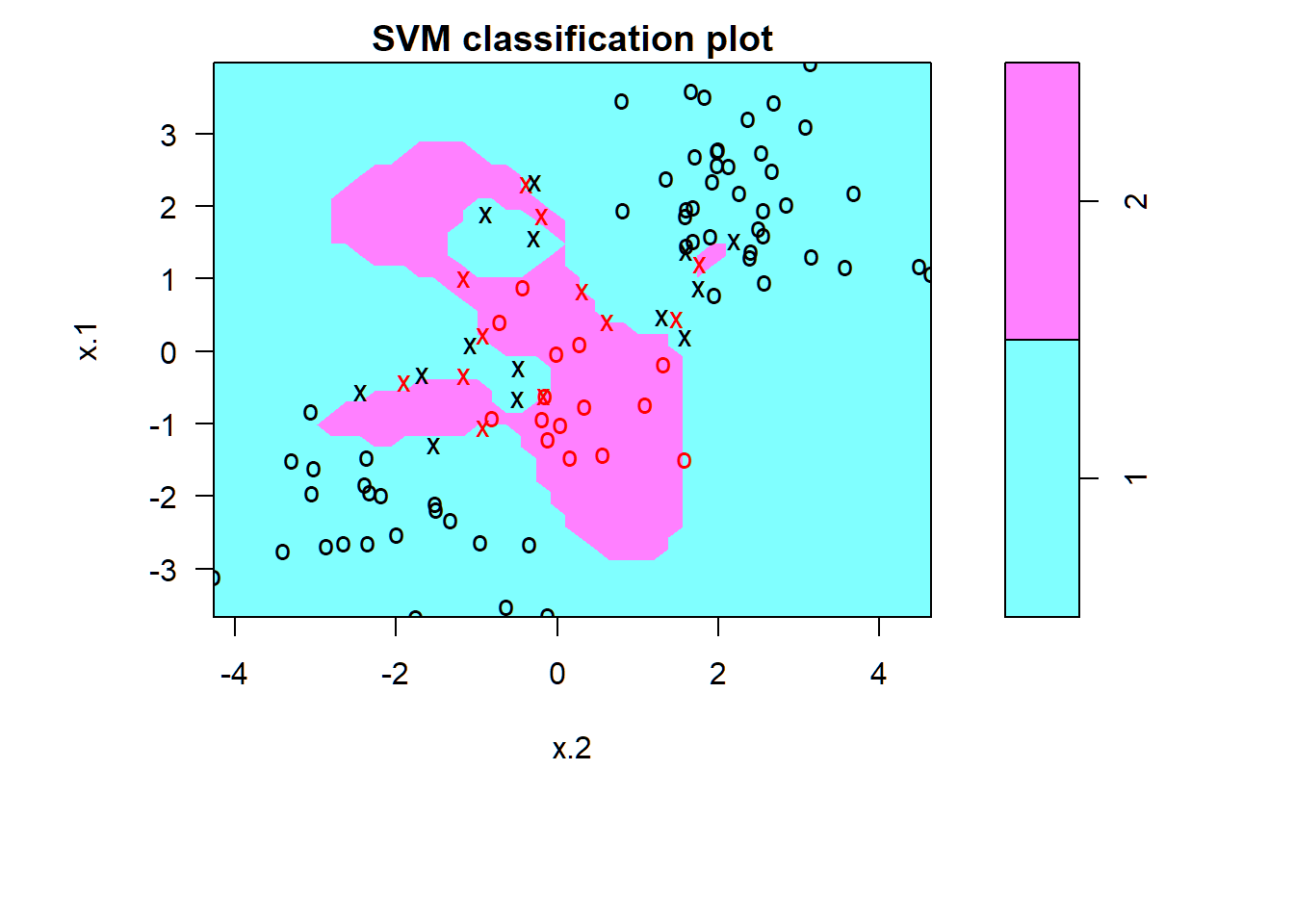
Cross-valiation on \(C,\gamma\)
Optimal parameter values determined by 10-fold cross-validation on training set: \(C=1,\gamma=2\)
> set.seed(1)
> tune.out=tune(svm, y~., data=dat[train,], kernel="radial",
+ ranges=list(cost=c(0.1,1,10,100,1000),
+ gamma=c(0.5,1,2,3,4)))
> summary(tune.out)
Parameter tuning of 'svm':
- sampling method: 10-fold cross validation
- best parameters:
cost gamma
1 2
- best performance: 0.12
- Detailed performance results:
cost gamma error dispersion
1 1e-01 0.5 0.27 0.11595018
2 1e+00 0.5 0.13 0.08232726
3 1e+01 0.5 0.15 0.07071068
4 1e+02 0.5 0.17 0.08232726
5 1e+03 0.5 0.21 0.09944289
6 1e-01 1.0 0.25 0.13540064
7 1e+00 1.0 0.13 0.08232726
8 1e+01 1.0 0.16 0.06992059
9 1e+02 1.0 0.20 0.09428090
10 1e+03 1.0 0.20 0.08164966
11 1e-01 2.0 0.25 0.12692955
12 1e+00 2.0 0.12 0.09189366
13 1e+01 2.0 0.17 0.09486833
14 1e+02 2.0 0.19 0.09944289
15 1e+03 2.0 0.20 0.09428090
16 1e-01 3.0 0.27 0.11595018
17 1e+00 3.0 0.13 0.09486833
18 1e+01 3.0 0.18 0.10327956
19 1e+02 3.0 0.21 0.08755950
20 1e+03 3.0 0.22 0.10327956
21 1e-01 4.0 0.27 0.11595018
22 1e+00 4.0 0.15 0.10801234
23 1e+01 4.0 0.18 0.11352924
24 1e+02 4.0 0.21 0.08755950
25 1e+03 4.0 0.24 0.10749677
> table(true=dat[-train,"y"],
+ pred=predict(tune.out$best.model,newdata=dat[-train,]))
pred
true 1 2
1 74 3
2 7 16ROC curve: function
Function to obtain ROC curve:
> library(ROCR)
> rocplot=function(pred, truth, ...){
+ predob = prediction(pred, truth)
+ perf = performance(predob, "tpr", "fpr")
+ plot(perf,...)}Fitted values and ROC curves
> # apply optimal SVM to training set for prediction
> svmfit.opt=svm(y~., data=dat[train,], kernel="radial",
+ gamma=2, cost=1,decision.values=T)
> fitted=attributes(predict(svmfit.opt,dat[train,],
+ decision.values=TRUE))$decision.values
> par(mfrow=c(1,2))
> rocplot(fitted,dat[train,"y"],main="Training Data")
> # another SVM built on training set and
> # applied to it for prediction with gamma=50
> svmfit.flex=svm(y~., data=dat[train,], kernel="radial",
+ gamma=50, cost=1, decision.values=T)
> fitted=attributes(predict(svmfit.flex,dat[train,],
+ decision.values=T))$decision.values
> rocplot(fitted,dat[train,"y"],add=T,col="red")decision.values: fitted value \(f_K(x; \hat{\alpha},\hat{\beta_0}) = \sum_{i=1}^n \hat{a}_i y_i K(x,x_i) + \hat{\beta}_0\) for a value \(x\) of feature vector \(X\)attributes(predict.obj)$decision.values: \(\hat{f}(x)\) when \(x\) is a test observation
Fitted values and ROC curves
> # optimal model SVM applied to test set for prediction
> fitted=attributes(predict(svmfit.opt,dat[-train,],
+ decision.values=T))$decision.values
> rocplot(fitted,dat[-train,"y"],main="Test Data")
> # non-optimal SVM applied to tet set for prediction
> fitted=attributes(predict(svmfit.flex,dat[-train,],
+ decision.values=T))$decision.values
> rocplot(fitted,dat[-train,"y"],add=T,col="red")- “optimal” \(\gamma=2\) obtained from range
gamma=c(0.5,1,2,3,4)by 10-fold cross-validation on training set; so, another value for \(\gamma\) may improve the “optimal” SVM on training set - another SVM is built with \(\gamma=50\) and \(C=1\) on training set
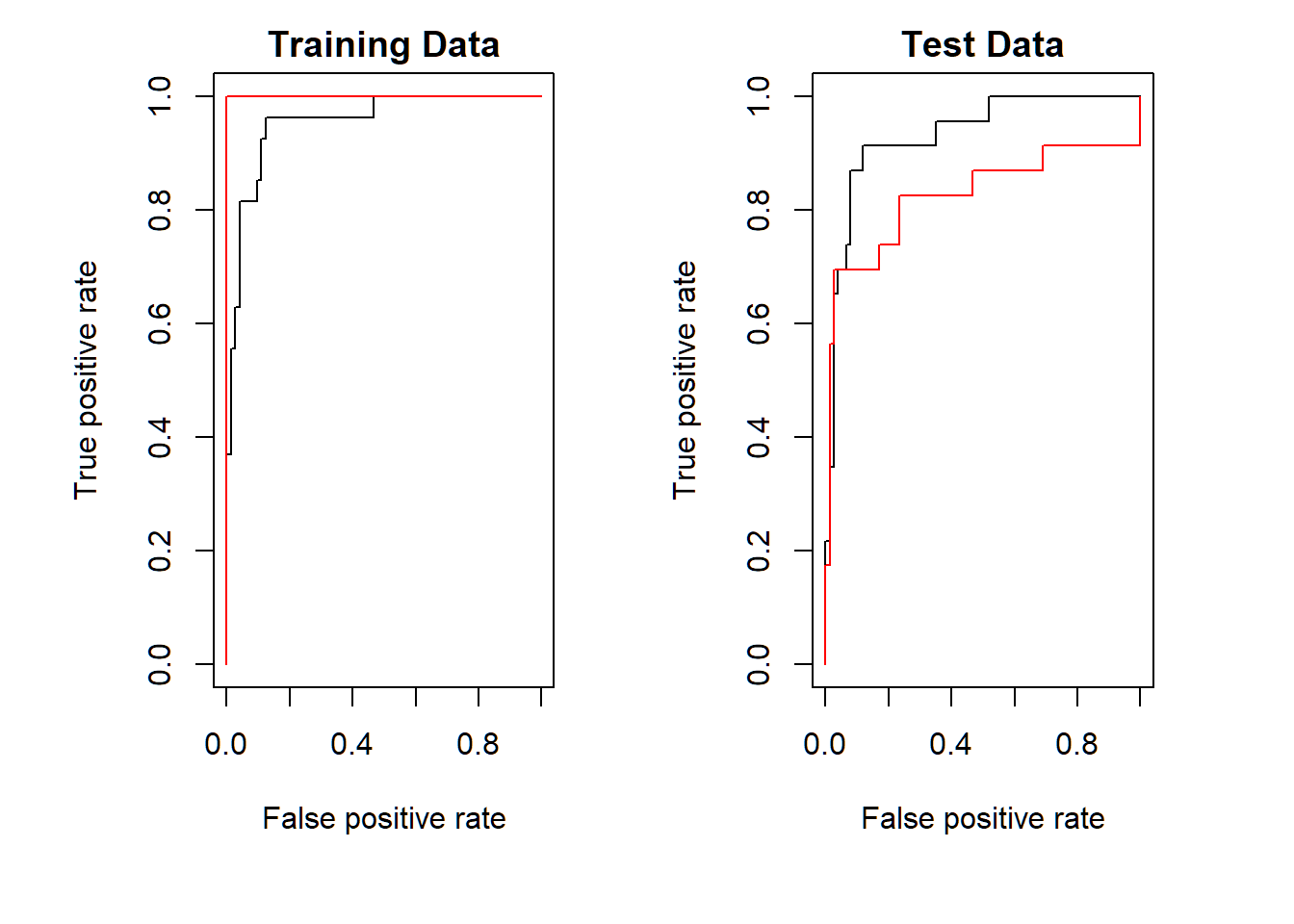
ROC curves: radial SVM
SVM with \(\gamma=50,C=1\) (red) more accurate than SVM with \(\gamma=2,C=1\) (black) on training set; the opposite happens on test set
Support vector machines (SVMs): Example 2
Generate data
- \(n=250\) observations on \(p=2\) features \(X_1,X_2\)
- Class \(1\): \(\quad 150\) observations; \[x_i \sim \text{Gaussian}(\mu_1=(2,2)^T,\mathbf{I}),i=1,\ldots,100\\ x_i \sim \text{Gaussian}(\mu_2=(-2,-2)^T,\mathbf{I}),i=101,\ldots,150 \]
- Class \(2\): \(\quad 50\) observations; \(x_i \sim \text{Gaussian}(\mu_3=(0,0)^T,\mathbf{I}),i=151,\ldots,200\)
- Class \(0\): \(\quad 50\) observations; \(x_i \sim \text{Gaussian}(\mu_4=(0,2)^T,\mathbf{I}),i=201,\ldots,250\)
> set.seed(1); x=matrix(rnorm(200*2),ncol=2)
> x[1:100,]=x[1:100,]+2; x[101:150,]=x[101:150,]-2
> y=c(rep(1,150),rep(2,50))
> x=rbind(x, matrix(rnorm(50*2), ncol=2))
> y=c(y, rep(0,50)); x[y==0,2]=x[y==0,2]+2
> dat=data.frame(x=x, y=as.factor(y))Scatter plot
3 classes with nonlinear class boundary:
> par(mar=c(5.5,4.5,1.2,.5),oma=c(1.3,1.3,0.3,1.3))
> plot(x,col=(y+1),xlab=expression(X[1]),ylab=expression(X[2]))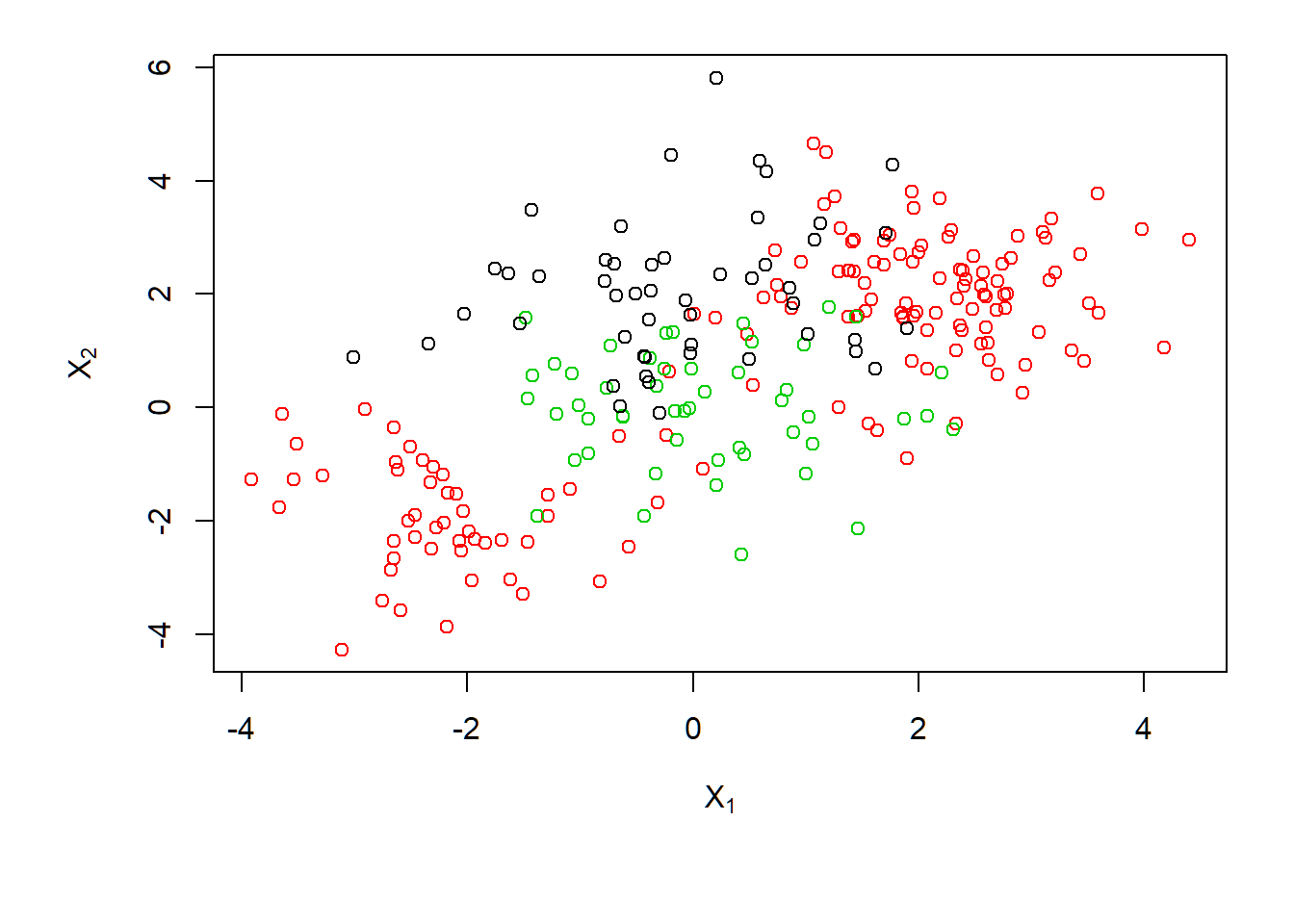
SVM with radial kernel
> svmfit=svm(y~., data=dat, kernel="radial", cost=10, gamma=1)
> plot(svmfit, dat)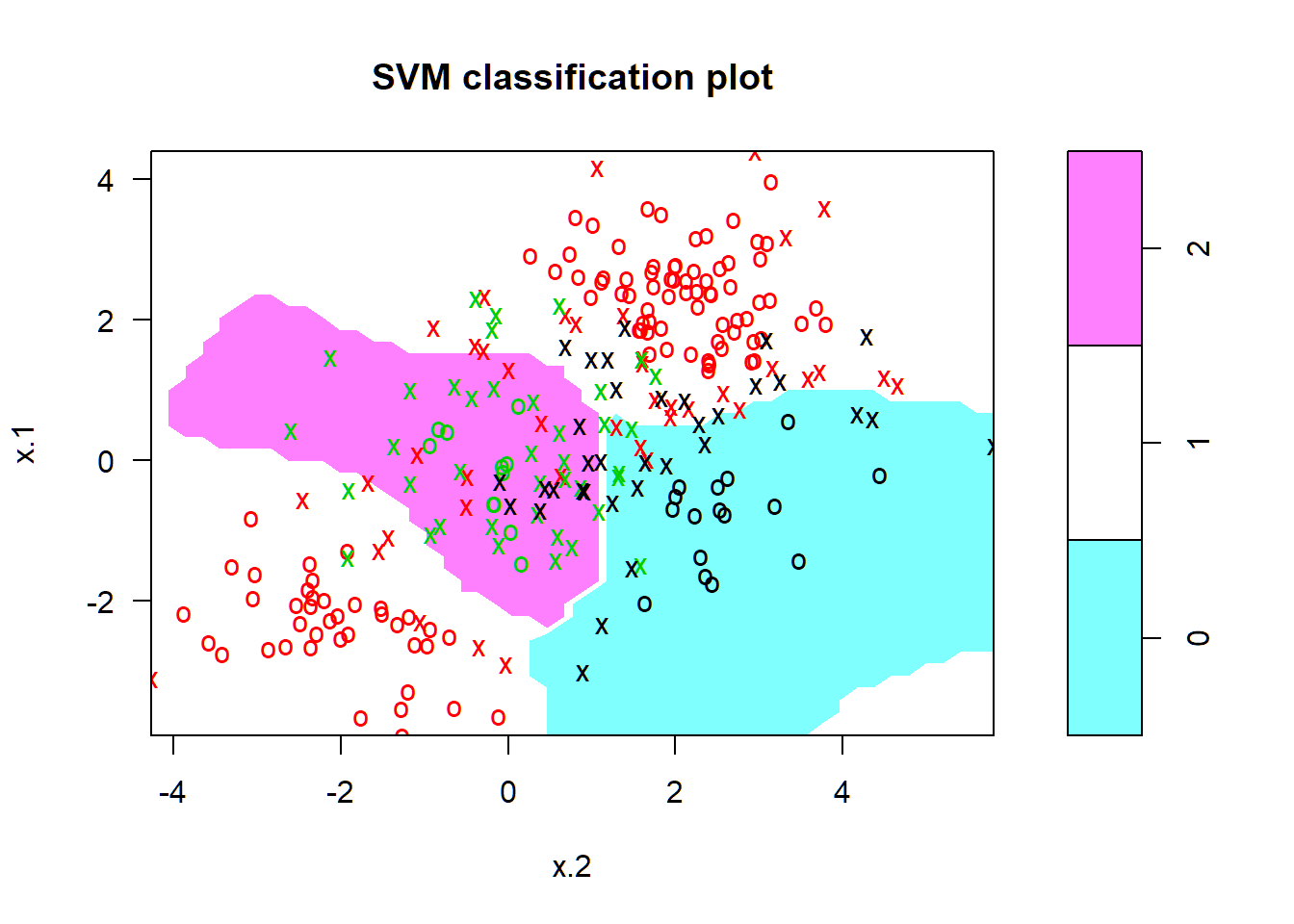
Support vector machines (SVMs): Example 3
Gene expression data
The dataset Khan{ISLR} consists of a number of tissue samples corresponding to four distinct types of small round blue cell tumors. For each tissue sample, there are 2308 gene expression measurements:
> library(ISLR)
> names(Khan);
[1] "xtrain" "xtest" "ytrain" "ytest"
> dim(Khan$xtrain) # training set x-pression matrix
[1] 63 2308
> dim(Khan$xtest) # test set x-pression matrix
[1] 20 2308
> length(Khan$ytrain) # training set class labels
[1] 63
> length(Khan$ytest) # test set class labels
[1] 20
> class(Khan$ytrain)
[1] "numeric"Khan$ytrainandKhan$ytest: both numeric
Information on classes
Number of observations for each tumor/caner type in training set and in test set:
> table(Khan$ytrain)
1 2 3 4
8 23 12 20
> table(Khan$ytest)
1 2 3 4
3 6 6 5
> dat=data.frame(x=Khan$xtrain, y=as.factor(Khan$ytrain))- Predict cancer type using gene expression measurements
- There are a very large number of features relative to the number of observations. So, it may be easier to find separating hyperplanes than using an SVM with radial kernel
SVM with linear kernel: training
SVM with linear kernel and \(C=10\) has 0 training error:
> out=svm(y~., data=dat, kernel="linear",cost=10)
> summary(out)
Call:
svm(formula = y ~ ., data = dat, kernel = "linear",
cost = 10)
Parameters:
SVM-Type: C-classification
SVM-Kernel: linear
cost: 10
gamma: 0.0004332756
Number of Support Vectors: 58
( 20 20 11 7 )
Number of Classes: 4
Levels:
1 2 3 4
> table(out$fitted, dat$y)
1 2 3 4
1 8 0 0 0
2 0 23 0 0
3 0 0 12 0
4 0 0 0 20SVM with linear kernel: prediction
SVM with linear kernel and \(C=10\) misclassified \(2\) observations in test set:
> dat.te=data.frame(x=Khan$xtest, y=as.factor(Khan$ytest))
> pred.te=predict(out, newdata=dat.te)
> table(pred.te, dat.te$y)
pred.te 1 2 3 4
1 3 0 0 0
2 0 6 2 0
3 0 0 4 0
4 0 0 0 5Support vector machines (SVMs): Example 4
Heart disease data Heart
- 13 features such as
Age,SexandChol(serum cholestoral in mg/dl), and \(303\) observations - Target: to predict whether an individual has heart disease, i.e.,
AHD=YesorAHD=No
> heartData=read.table("Heart.txt", sep=",",header=TRUE)
> heartData$X=NULL; heartData=na.omit(heartData)
> dim(heartData)
[1] 297 14
> heartData[1:3,c(1,2,5,14)]
Age Sex Chol AHD
1 63 1 233 No
2 67 1 286 Yes
3 67 1 229 Yes- number of features not too small or large relative to number of observations
SVM: linear kernel
SVM with linear kernel and optimal cost \(C=0.1\) misclassified \(14\) observations in test set:
> set.seed(123)
> trainId=sample(1:nrow(heartData),floor(0.7*nrow(heartData)))
> library(e1071)
> svmLKTune=tune(svm,AHD~., data=heartData[trainId,],
+ kernel="linear",
+ ranges=list(cost=c(0.1,1,10,100,1000)))
> predLK=predict(svmLKTune$best.model,heartData[-trainId,])
> table(predicted=predLK, truth=heartData[-trainId,]$AHD)
truth
predicted No Yes
No 44 8
Yes 6 32- when number of features \(p\) is not much larger than sample size \(n\), hyperplanes may not do a good job in separating observations into their classes
SVM: radial kernel
SVM with radial kernel and optimal \(C=10,\gamma=0.5\) misclassified \(27\) observations in test set:
> SVMRKTune=tune(svm, AHD~., data=heartData[trainId,],
+ kernel="radial",
+ ranges=list(cost=c(0.1,1,10,100,1000),
+ gamma=c(0.5,1,2,3,4,5,6)))
> predRK=predict(SVMRKTune$best.model,heartData[-trainId,])
> table(predicted=predRK, truth=heartData[-trainId,]$AHD)
truth
predicted No Yes
No 32 9
Yes 18 31- an optimal model selected by cross-validation depends on quality of training set, number of folds used, and ranges for tuning parameters
- optimal radial SVM not necessarily better than optimal linear SVM
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19041)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] stringr_1.3.1 xfun_0.4 digest_0.6.18
[13] evaluate_0.12