Stat 437 Lecture Notes 8b
Xiongzhi Chen
Washington State University
Principal component analysis (PCA): software implementation
R software and commands
R libraries and commands needed:
- R built-in command
prcomp{stats}to implement PCA - R built-in command
biplot.princomp{stats}to create biplot for PCA
Command prcomp
prcomp{stats}performs PCA on a data matrix and returns the results as an object of classprcompThe calculation is done by a singular value decomposition (SVD) of the (centered and possibly scaled) data matrix but not by using the command
eigenon the covariance matrix. This is generally the preferred method for numerical accuracy. Theprintmethod for these objects prints the results in a nice format, and theplotmethod produces a scree plot
Command prcomp
The basic syntax of prcomp{stats} is:
prcomp(x, retx = TRUE, center = TRUE, scale. = FALSE,
tol = NULL, rank. = NULL, ...)x: a numeric or complex matrix (or data frame) which provides the data for PCA.retx: a logical value indicating whether the rotated variables (i.e., score vectors) should be returned.center: a logical value indicating whether the variables (i.e., features) should be shifted to be zero centered. Alternately, a vector of length equal the number of columns ofxcan be supplied. The value is passed toscale.
Command prcomp
Basic syntax:
prcomp(x, retx = TRUE, center = TRUE, scale. = FALSE,
tol = NULL, rank. = NULL, ...)scale.: a logical value indicating whether the variables should be scaled to have unit variance before the analysis takes place. The default isFALSEbut in general scaling is advisable. Alternatively, a vector of length equal the number of columns ofxcan be supplied. The value is passed toscale.
Command prcomp
Basic syntax:
prcomp(x, retx = TRUE, center = TRUE, scale. = FALSE,
tol = NULL, rank. = NULL, ...)tol: a value indicating the magnitude below which principal components (PCs) should be omitted. PCs are omitted if their standard deviations (i.e., if the Euclidean norms of their score vectors) are less than or equal totoltimes the standard deviation of the first PC. The default isnulland no components are omitted (unlessrank.is specified less thanmin(dim(x)). Also,tol=0ortol=sqrt(.Machine$double.eps)can be set, which would omit essentially constant PCs.
Command prcomp
Basic syntax:
prcomp(x, retx = TRUE, center = TRUE, scale. = FALSE,
tol = NULL, rank. = NULL, ...)rank.: a number specifying the maximal rank, i.e., maximal number of principal components to be used. It can be set as an alternative to or in addition totol, and is useful notably when the desired rank is considerably smaller than the dimensions of the matrixx.
Command prcomp
prcomp returns a list with class “prcomp” containing the following components:
sdev: a vector of standard deviations of PCs; the \(i\)-th entry ofsdevis for PC \(i\). So, whenxis column-centered,(nrow(x)-1)*(sdev^2)are the eigenvalues oft(x)%*%x, and their sum is the total variability for data in the column-centeredx.rotation: matrix of loading vectors, i.e., the matrix whose columns are eigenvectors oft(x)%*%xwhenxis column-centered (and column-scaled).
Command prcomp
prcomp returns a list with class “prcomp” containing the following components:
x: ifretx=TRUE, then the value of the rotated data is returned, i.e., ifretx=TRUE,xis the centered (and scaled if requested) data multiplied by therotationmatrix, i.e., ifretx=TRUE,xis the matrix whose columns are score vectors. The \(i\)-th column ofxpairs with the \(i\)-th column ofrotation, and both are for PC \(i\).centerandscale: the centering and scaling used ifcenter=TRUEandscale.=TRUE; otherwise, either or both areFALSE.
Command biplot.princomp
biplot.princomp{stats} (i.e., biplot{stats} when applied to an object of class “prcomp” ) produces a biplot from the output of command prcomp (or princomp). Its basic syntax is:
biplot(x,choices = 1:2,scale = 1,pc.biplot=FALSE,cex=1,...)x: an object of class “prcomp” (or “princomp” whose details are given in Appendix 1).choices: a length 2 vector specifying the principal components to plot.cex: font size if biplot is annotated by labels of observations.
Command biplot.princomp
Basic syntax:
biplot(x,choices = 1:2,scale = 1,pc.biplot=FALSE,cex=1,...)scale: variables are scaled bylambda^scaleand observations bylambda^(1-scale), wherelambdaare the singular values as computed byprincomp. Normally0 <= scale <= 1, and a warning will be issued if the specified scale is outside this range. By default,scale=1.pc.biplot: iftrue, use what Gabriel (1971) refers to as a “principal component biplot”, withlambda=1and observations scaled up bysqrt(n)and variables scaled down bysqrt(n).
Principal component analysis: Example 1
US arrests data
The USArrests data set (included in the library ISLR or ggplot2) contains arrests per 100,000 residents in each of the 50 US states in 1973. There are
- 4 features: 3 types of crimes
Murder,Assault,Rape, andUrbanPop(percent of population living in urban areas)
> library(ISLR); dim(USArrests)
[1] 50 4
> head(USArrests)
Murder Assault UrbanPop Rape
Alabama 13.2 236 58 21.2
Alaska 10.0 263 48 44.5
Arizona 8.1 294 80 31.0
Arkansas 8.8 190 50 19.5
California 9.0 276 91 40.6
Colorado 7.9 204 78 38.7Correlations among features
addCoef.col="grey": add correlation coefficient with color;tl.cex=0.57: font size for variables
> corMatFeature = cor(USArrests); library(corrplot)
> corrplot(corMatFeature,method="shade",addCoef.col="grey",
+ tl.cex=0.57, mar = c(4.5,1,1,1))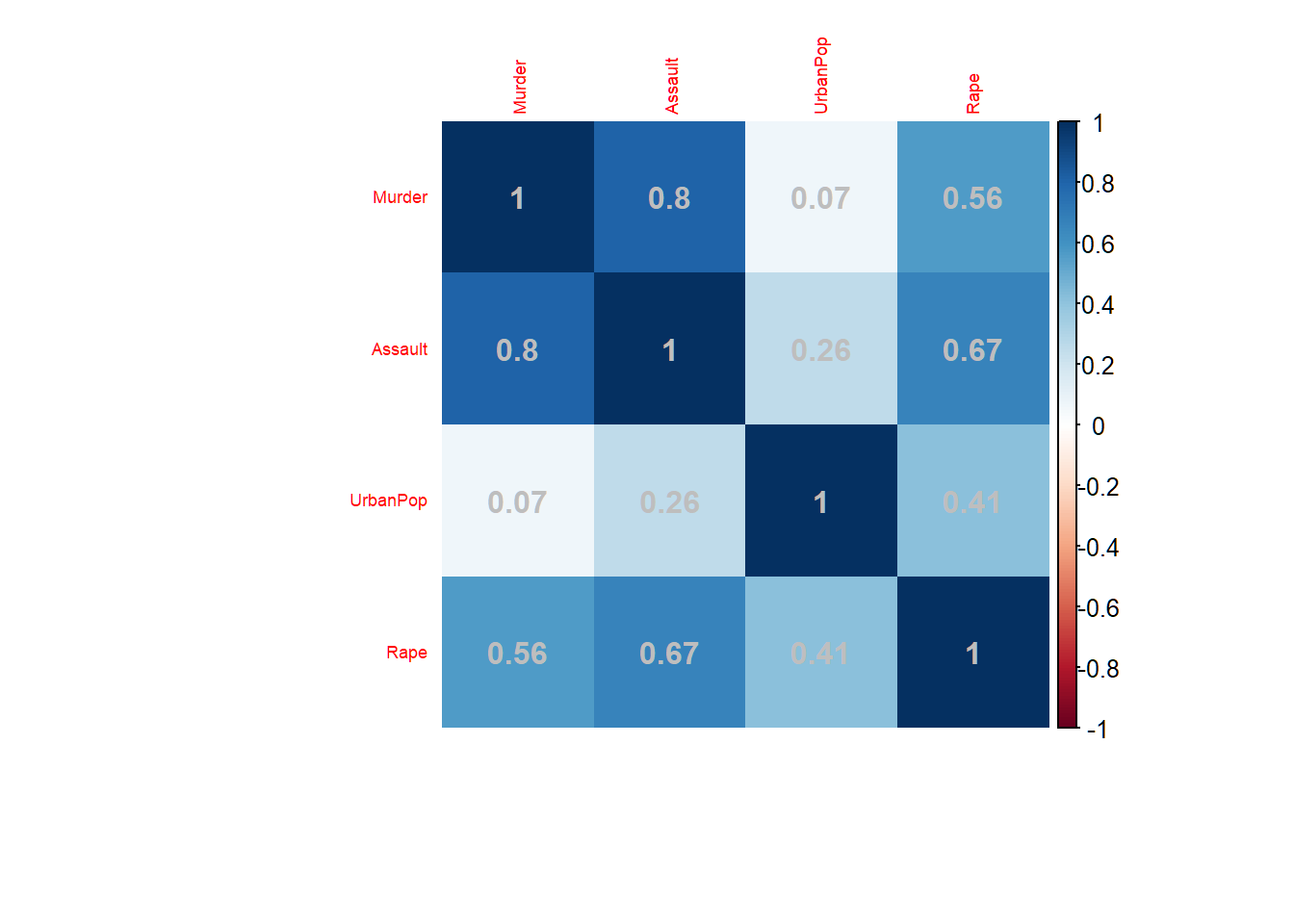
Dendrogram
> library(ggdendro); library(ggplot2)
> Xsd=scale(USArrests,center=TRUE,scale=TRUE)
> hc=hclust(dist(Xsd),"ave"); source("C:\\MathRepo\\stat437Online\\Plotggdendro.r")
> plot_ggdendro(dendro_data_k(hc,5),direction ="tb",expand.y = 0.2)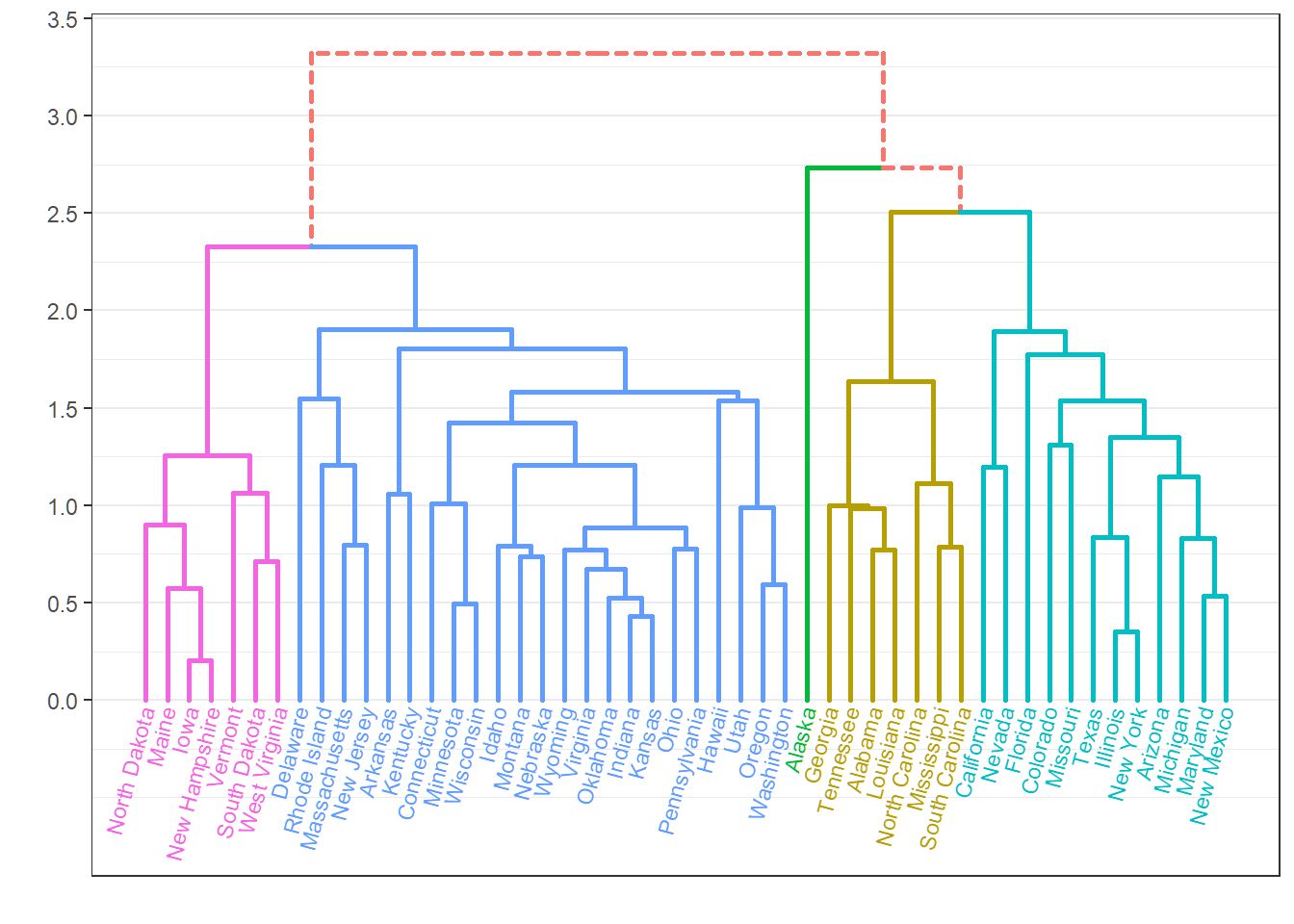
PCA of standardized data
- Acronym “PC” means “loading vector”
> # standardize data
> Xsd = scale(USArrests,center=TRUE,scale=TRUE)
> # total variability of standardized data
> sum(Xsd^2)
[1] 196
> # PCA for standardized data
> pcaXsd = prcomp(Xsd)
> # directions or loading vectors
> pcaXsd$rotation
PC1 PC2 PC3 PC4
Murder -0.5358995 0.4181809 -0.3412327 0.64922780
Assault -0.5831836 0.1879856 -0.2681484 -0.74340748
UrbanPop -0.2781909 -0.8728062 -0.3780158 0.13387773
Rape -0.5434321 -0.1673186 0.8177779 0.08902432
> # successive maximal variabilities of projections
> (nrow(USArrests)-1)*((pcaXsd$sdev)^2)
[1] 121.531837 48.498492 17.471596 8.498074
> # cumulative proportion of variance explained
> cumsum((nrow(USArrests)-1)*((pcaXsd$sdev)^2))/sum(Xsd^2)
[1] 0.6200604 0.8675017 0.9566425 1.0000000- Note
(nrow(USArrests)-1)*((pcaXsd$sdev)^2)
Score matrix
- Score matrix is \(n \times p\); each column is a score vector
- Each row is labelled by observation label
> options(digits=5)
> pcaXsd$x
PC1 PC2 PC3 PC4
Alabama -0.975660 1.122001 -0.439804 0.1546966
Alaska -1.930538 1.062427 2.019500 -0.4341755
Arizona -1.745443 -0.738460 0.054230 -0.8262642
Arkansas 0.139999 1.108542 0.113422 -0.1809736
California -2.498613 -1.527427 0.592541 -0.3385592
Colorado -1.499341 -0.977630 1.084002 0.0014502
Connecticut 1.344992 -1.077984 -0.636793 -0.1172787
Delaware -0.047230 -0.322089 -0.711410 -0.8731133
Florida -2.982760 0.038834 -0.571032 -0.0953170
Georgia -1.622807 1.266088 -0.339018 1.0659745
Hawaii 0.903484 -1.554676 0.050272 0.8937332
Idaho 1.623319 0.208853 0.257190 -0.4940879
Illinois -1.365052 -0.674988 -0.670686 -0.1207949
Indiana 0.500381 -0.150039 0.225763 0.4203976
Iowa 2.230996 -0.103008 0.162910 0.0173795
Kansas 0.788872 -0.267449 0.025296 0.2044210
Kentucky 0.743313 0.948807 -0.028084 0.6638172
Louisiana -1.549091 0.862300 -0.775606 0.4501578
Maine 2.372740 0.372609 -0.065022 -0.3271385
Maryland -1.745647 0.423357 -0.155670 -0.5534506
Massachusetts 0.481280 -1.459677 -0.603372 -0.1777939
Michigan -2.087250 -0.153835 0.381000 0.1013431
Minnesota 1.675670 -0.625907 0.151532 0.0666403
Mississippi -0.986479 2.369737 -0.733363 0.2133420
Missouri -0.689784 -0.260708 0.373650 0.2235548
Montana 1.173538 0.531479 0.244408 0.1224986
Nebraska 1.252916 -0.192004 0.173809 0.0157332
Nevada -2.845505 -0.767805 1.151688 0.3113544
New Hampshire 2.359956 -0.017901 0.036485 -0.0328043
New Jersey -0.179741 -1.434937 -0.756770 0.2409366
New Mexico -1.960124 0.141413 0.181846 -0.3361211
New York -1.665667 -0.814911 -0.636612 -0.0133488
North Carolina -1.112088 2.205611 -0.854892 -0.9447896
North Dakota 2.962152 0.593097 0.298249 -0.2514346
Ohio 0.223694 -0.734778 -0.030826 0.4691528
Oklahoma 0.308649 -0.284961 -0.015156 0.0102285
Oregon -0.058528 -0.535970 0.930387 -0.2353909
Pennsylvania 0.879487 -0.565361 -0.396602 0.3554524
Rhode Island 0.855091 -1.476983 -1.356177 -0.6074027
South Carolina -1.307450 1.913973 -0.297517 -0.1301454
South Dakota 1.967797 0.815068 0.385381 -0.1084705
Tennessee -0.989694 0.851605 0.186193 0.6463027
Texas -1.341518 -0.408335 -0.487123 0.6367311
Utah 0.545032 -1.456715 0.290776 -0.0814867
Vermont 2.773256 1.388194 0.832808 -0.1434337
Virginia 0.095367 0.197728 0.011595 0.2092464
Washington 0.214723 -0.960374 0.618591 -0.2186282
West Virginia 2.087393 1.410526 0.103722 0.1305831
Wisconsin 2.058812 -0.605125 -0.137469 0.1822534
Wyoming 0.623101 0.317787 -0.238240 -0.1649769Biplot for PCA
> # use reflection to comply with Textbook
> pcaXsd$rotation = -pcaXsd$rotation; pcaXsd$x = -pcaXsd$x
> par(mar=c(7,4.5,1.5,1),mgp=c(1.5,0.5,0))
> biplot(pcaXsd,choices=1:2,scale=1,pc.biplot=T,cex=0.5)
> # `cex=0.5`: font size for text labels
> abline(v=0, lty=2, col="grey50")
> abline(h=0, lty=2, col="grey50")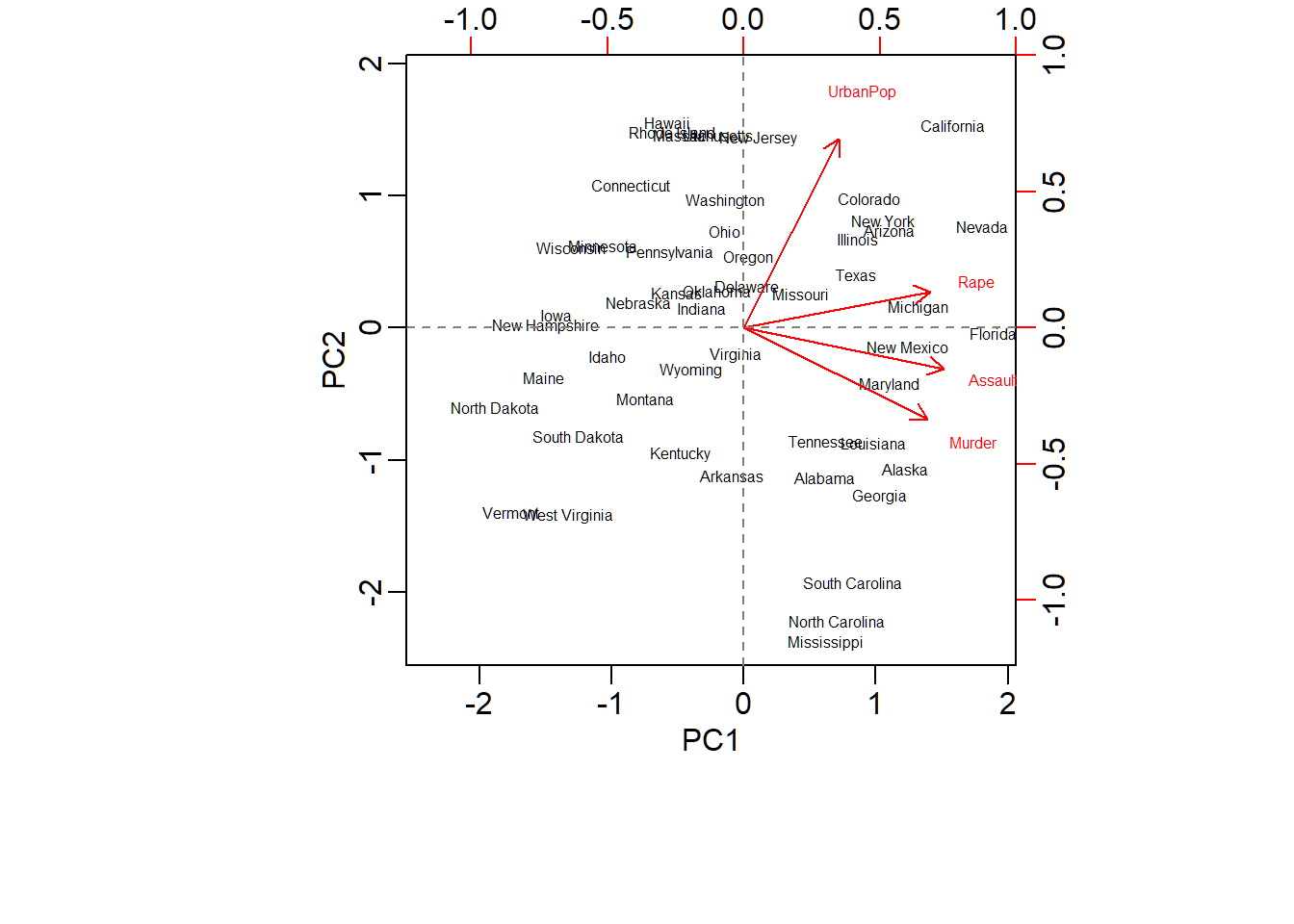
Customized biplot
Cluster memberships of US states superimposed on biplot of PCA of USArrests data
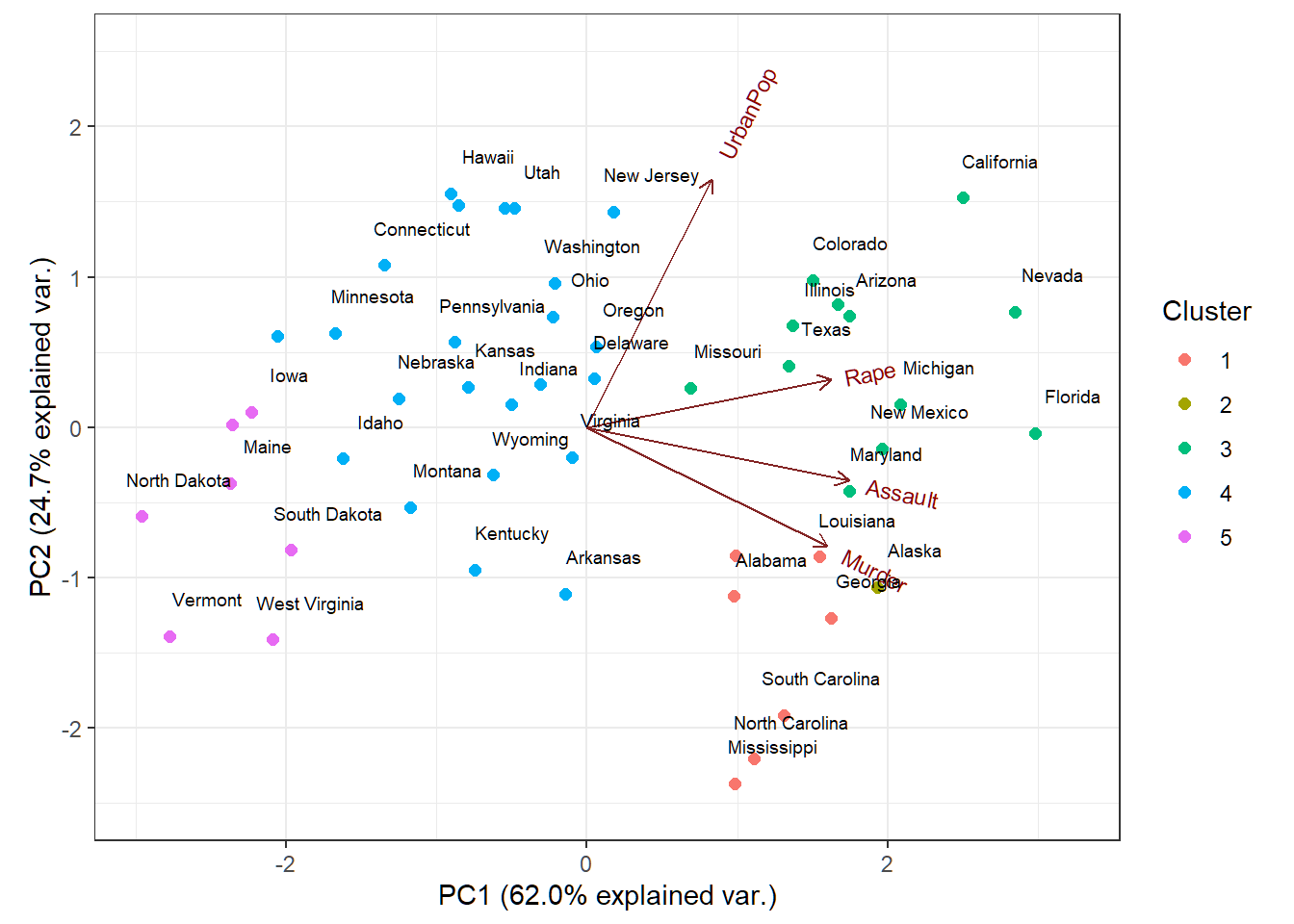
Remark on prcomp
Let \(\tilde{\mathbf{X}} \in \mathbb{R}^{n \times p}\) be the standardized data matrix that is obtained from the data matrix \({\mathbf{X}} \in \mathbb{R}^{n \times p}\)
The
prcomp$sdevare square roots of the eigenvalues of the matrix \(\hat{\mathbf{X}}^T\hat{\mathbf{X}}\) with \[ \hat{\mathbf{X}}=(n-1)^{-1/2}\tilde{\mathbf{X}}, \] such that \(\text{tr}(\hat{\mathbf{X}}^T\hat{\mathbf{X}})=p\)\(\tilde{\mathbf{X}}^T\tilde{\mathbf{X}}\) and \(\hat{\mathbf{X}}^T\hat{\mathbf{X}}\) have the same set of orthonormal eigenvectors (modulo \(\pm\) signs ) but the eigenvalues of \(\tilde{\mathbf{X}}^T\tilde{\mathbf{X}}\) are \(n-1\) times those of \(\hat{\mathbf{X}}^T\hat{\mathbf{X}}\) when both sets of eigenvalues are arranged in descending order
Principal component analysis: Example 2
Human cancer microarray data
Human cancer data:
- 6830 genes (i.e., \(p=6830\) features) and their expressions
- 64 samples (i.e., \(64\) cancer cell lines)
- 14 different cancer types; information on cancer types at wiki-NCI60
We will pick observations for 3 caner types “BREAST”, “OVARIAN” and “LEUKEMIA”, and this gives the uncentered data matrix \(\mathbf{X} \in \mathbb{R}^{19 \times 6830}\) (i.e., \(p=6830\) features and \(n=19\) observations)
PCA on standardized data
> library(ISLR);
> nci.labs = NCI60$labs # cancer types or cell lines
> nci.data = NCI60$data # expression data matrix
> # pick 3 cancer types: BREAST, OVARIAN, LEUKEMIA
> nci.labs1 = nci.labs[nci.labs %in%
+ c('BREAST', 'OVARIAN', 'LEUKEMIA')]
> nci.data1 = nci.data[nci.labs %in%
+ c('BREAST', 'OVARIAN', 'LEUKEMIA'),]
> # standardize data
> Xsd = scale(nci.data1,center=TRUE,scale=TRUE)
> # PCA for standardized data
> pcaXsd = prcomp(Xsd)
> names(pcaXsd)
[1] "sdev" "rotation" "center" "scale" "x"
> dim(nci.data) # dimensions of data matrix
[1] 64 6830
> dim(pcaXsd$rotation) # dimensions of loading matrix
[1] 6830 19
> dim(pcaXsd$x) # dimensions of score matrix
[1] 19 19Remarks on rank.
- In previous slide, \(19\) PCs, \(19\) loading vectors and \(19\) score vectors were produced because we used
tol=NULLandrank.=NULLin commandprcomp, which removed no PCs However, the \(19\)-th PC and its loading vector and score vector theoretically should not exist at all (since PCA was applied to the standardized data matrix \(\tilde{\mathbf{X}}\) which has rank \(18=\text{rank}(\tilde{\mathbf{X}})\)) but are by-products of machine precision. We can use
rank.=18to remove them.Since we did not remove any PC, the \(19\) PCs and their loading and score vectors, their PVEs and CPVE will appear in later slides (as illustrations for instruction)
Remarks on rank.
In practice, we need to specify argument
rank.to be \(\text{rank}(\tilde{\mathbf{X}})\) (i.e., the rank of \(\tilde{\mathbf{X}}\)) for commandprcompFor the standardized data matrix \(\tilde{\mathbf{X}} \in \mathbb{R}^{n \times p}\) that contains \(n\) linearly independent observations on \(p\) linearly independent features (,which is usually the case), its rank \[\text{rank}(\tilde{\mathbf{X}}) = \min\{n-1,p\}.\] Namely, for such \(\tilde{\mathbf{X}}\), we set
rank.to be \(\min\{n-1,p\}\) forprcomp
Obtain \(r=\text{rank}(\tilde{\mathbf{X}})\) PCs
- Standardized data matrix \(\tilde{\mathbf{X}}\) has rank \(r=18\), allowing for a maximum of \(18\) PCs when PCA is applied to \(\tilde{\mathbf{X}}\):
> library(ISLR); nci.labs = NCI60$labs; nci.data = NCI60$data
> nci.labs1 = nci.labs[nci.labs %in%
+ c('BREAST', 'OVARIAN', 'LEUKEMIA')]
> nci.data1 = nci.data[nci.labs %in%
+ c('BREAST', 'OVARIAN', 'LEUKEMIA'),]
> Xsd = scale(nci.data1,center=TRUE,scale=TRUE)
> # PCA for standardized data
> pcaXsd1 = prcomp(Xsd,rank.=nrow(nci.data1)-1)
> dim(nci.data) # dimensions of data matrix
[1] 64 6830
> dim(pcaXsd1$rotation) # dimensions of loading matrix
[1] 6830 18
> dim(pcaXsd1$x) # dimensions of score matrix
[1] 19 18
> cumsum(100*(nrow(nci.data1)-1)*pcaXsd1$sdev^2)/sum(Xsd^2) # CPVE
[1] 16.95366 27.33261 36.83971 44.15321 50.83002
[6] 56.55363 62.01891 66.77877 71.51763 75.68344
[11] 79.67929 83.60823 87.38751 90.98552 94.09808
[16] 96.83802 99.13764 100.00000 100.00000First 4 PC loading vectors
Create a histogram for entries of each of first 4 PC loading vectors:
> library(ggplot2); library(reshape2)
> # column names of loading matrix
> colnames(pcaXsd$rotation)
[1] "PC1" "PC2" "PC3" "PC4" "PC5" "PC6" "PC7" "PC8"
[9] "PC9" "PC10" "PC11" "PC12" "PC13" "PC14" "PC15" "PC16"
[17] "PC17" "PC18" "PC19"
> # store first 4 PC loading vectors in a matrix
> FourLD = pcaXsd$rotation[,1:4]
> # change column names of `FourLD`
> colnames(FourLD)=paste(colnames(FourLD),"loadings",sep=" ")
> colnames(FourLD)
[1] "PC1 loadings" "PC2 loadings" "PC3 loadings" "PC4 loadings"
> # melt the matrix `FourLD` for plotting
> dstack <- melt(FourLD)
> # create a histogram for entries of each loading vector
> lvPlot=ggplot(dstack,aes(x = value))+
+ geom_histogram(bins = 25)+
+ facet_wrap(~Var2,scales = "free_x")+theme_bw()First 4 PC loading vectors
Each loading vector has a lot of nonzero entries; there seem to be no dominant features (i.e., genes) in any direction
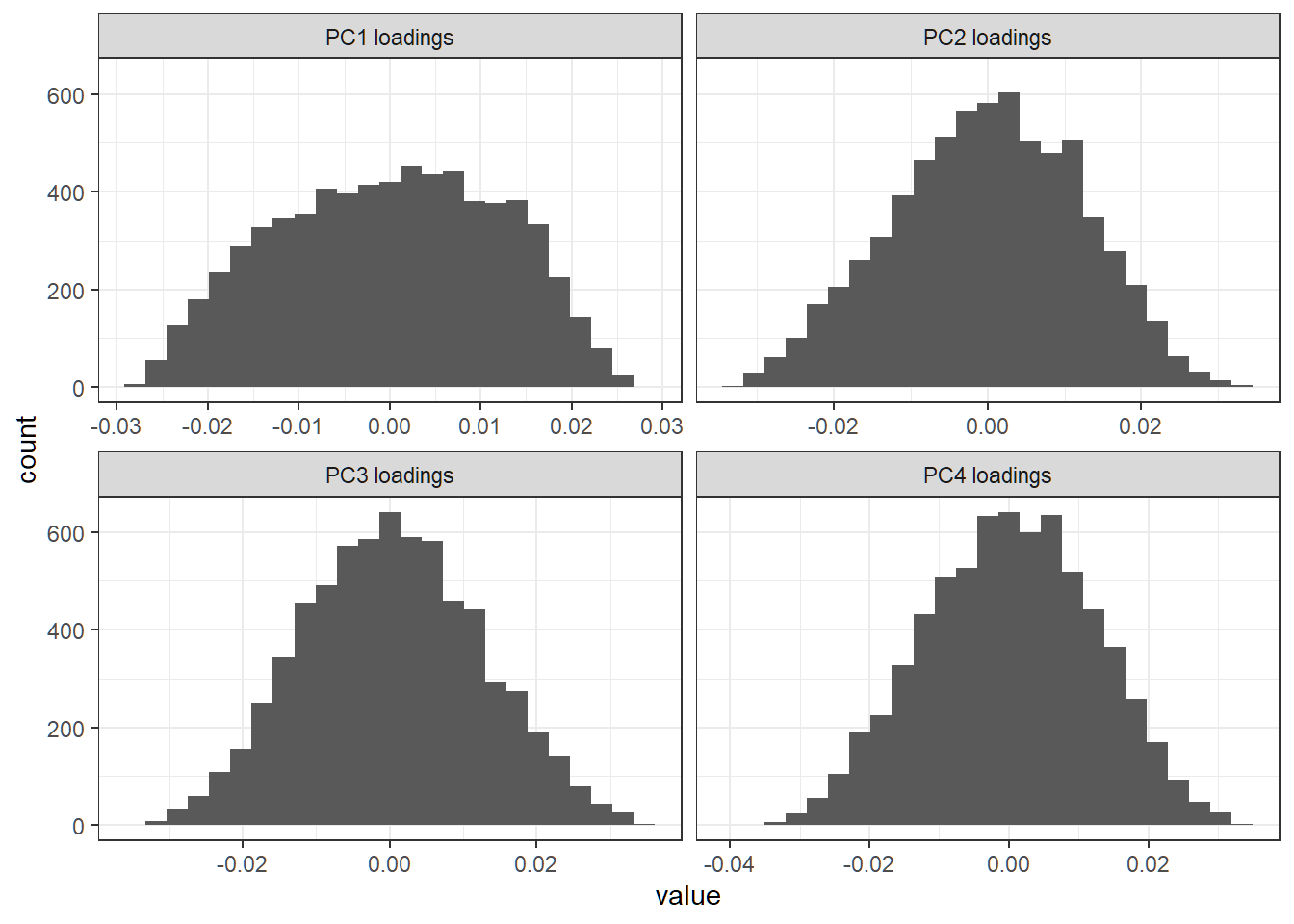
Proportion of variance explained
> # compute PVE for each PC; scaling (n-1) cancelled in ratio
> pve = 100*pcaXsd$sdev^2/sum(pcaXsd$sdev^2)
> pve
[1] 1.695366e+01 1.037896e+01 9.507100e+00 7.313498e+00
[5] 6.676814e+00 5.723603e+00 5.465282e+00 4.759858e+00
[9] 4.738859e+00 4.165811e+00 3.995856e+00 3.928936e+00
[13] 3.779277e+00 3.598014e+00 3.112560e+00 2.739935e+00
[17] 2.299622e+00 8.623618e-01 2.256130e-29
> # compute cumulative PVE (CPVE)
> cumsum(pve)
[1] 16.95366 27.33261 36.83971 44.15321 50.83002
[6] 56.55363 62.01891 66.77877 71.51763 75.68344
[11] 79.67929 83.60823 87.38751 90.98552 94.09808
[16] 96.83802 99.13764 100.00000 100.00000- The last, i.e., \(19\)-th, PVE should be theoretically \(0\) (since the standardized data matrix has rank \(18\)) but is numerically nonzero as
2.256130e-29due to machine precision. However, it is essentially \(0\)
Plot of PVE and CPVE
> par(mfrow=c(1,2),mar=c(9,4,1,3),mgp=c(1.8,0.6,0.3))
> plot(1:length(pve), pve,type="o",ylab="PVE",
+ xlab="Principal component",col="blue")
> plot(cumsum(pve),type="o",ylab="Cumulative PVE",
+ xlab="Number of first PCs",col="blue")
> abline(h = cumsum(pve)[11], col="red", lty=2)
> abline(v = 11, col="red", lty=2)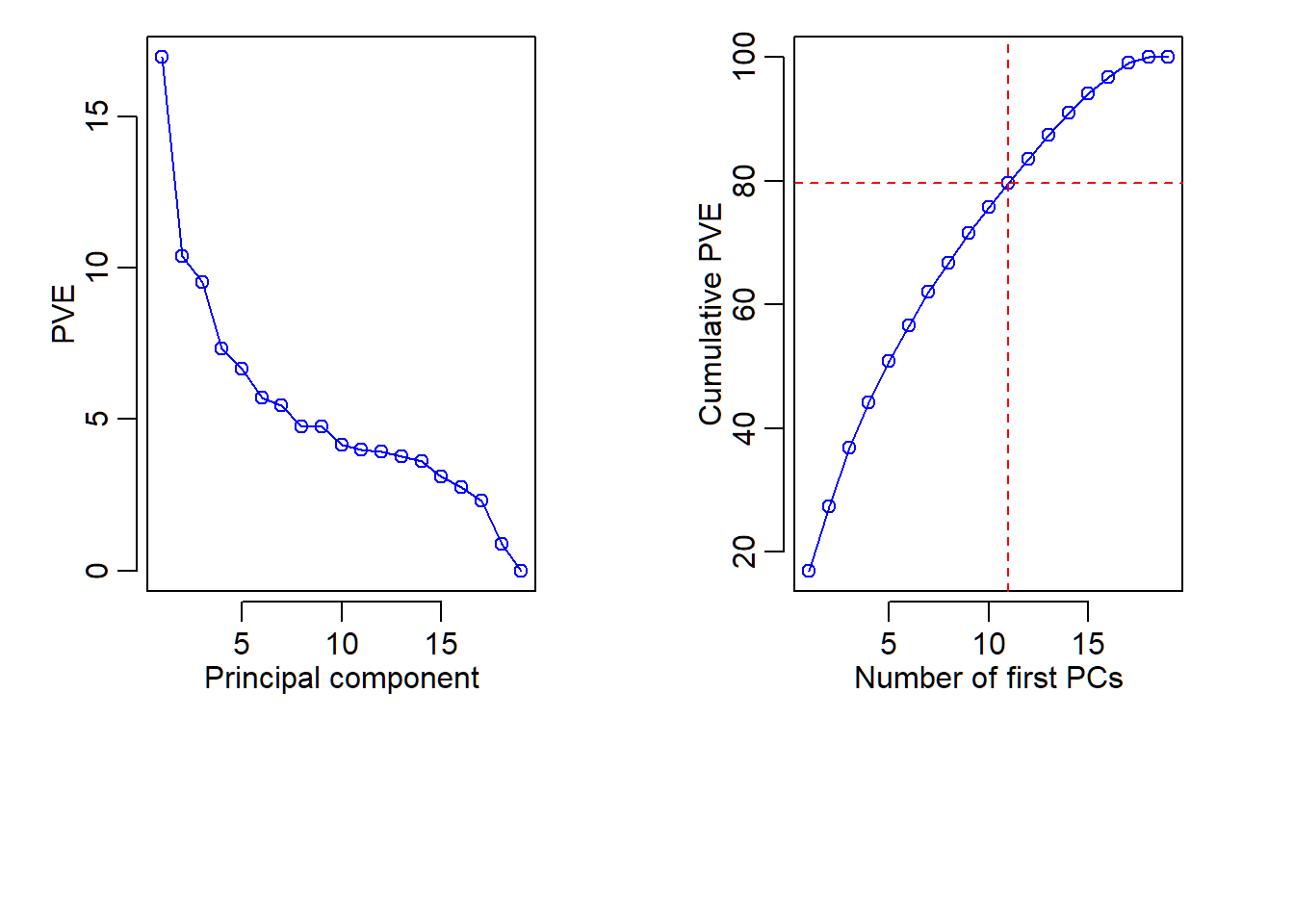
First 2 PC score vectors
- Data matrix does not have row names as cancer types
- Add rowname for row \(i\) of the score matrix (each of whose columns is a score vector) as the label (i.e., cancer type) for observation \(i\); “PC\(j\)” here means “score vector for PC\(j\)”
> scoresMat=as.data.frame(pcaXsd$x[,1:2])
> sMTmp=t(scoresMat); colnames(sMTmp)=nci.labs1
> scoresMat=t(sMTmp); scoresMat
PC1 PC2
BREAST -60.098790 1.588815
BREAST -31.889651 10.877669
BREAST -29.035831 6.582595
OVARIAN -19.721568 27.641286
OVARIAN -12.608223 14.464804
OVARIAN -26.962732 12.273994
OVARIAN -6.453167 9.465530
OVARIAN -22.091467 32.942808
OVARIAN -16.828185 19.572828
LEUKEMIA 17.967827 -20.708967
LEUKEMIA 40.596818 -8.642872
LEUKEMIA 48.904273 -5.795735
LEUKEMIA 66.947137 15.148187
LEUKEMIA 56.697268 15.532885
LEUKEMIA 23.671867 -8.962769
BREAST 9.829156 4.542899
BREAST -6.791608 5.910896
BREAST -17.605215 -66.753270
BREAST -14.527909 -65.681584Plot first 2 PC score vectors
> # store first 2 score vectors in a data.frame
> scoresMat=as.data.frame(pcaXsd$x[,1:2])
> # add cancer types for observations to be used as labels
> # when plotting scores
> scoresMat$Cancer=nci.labs1
> # create plot with `color=Cancer,shape=Cancer`
> library(ggplot2)
> pSV = ggplot(scoresMat,aes(PC1,PC2,
+ color=Cancer,shape=Cancer))+
+ geom_point()+theme_bw()+xlab("PC1 scores")+
+ ylab("PC2 scores")+
+ geom_hline(yintercept = 0,linetype="dotted")+
+ geom_vline(xintercept = 0,linetype="dotted")+
+ geom_text(aes(label=Cancer),hjust=-0.2, vjust=0.9,size=2)
> # geom_hline and geom_vline as center lines as referencesPlot first 2 PC score vectors
PC1 scores more indicative of “LEUKEMIA”, and PC2 scores of “OVARIAN”; other patterns may exist
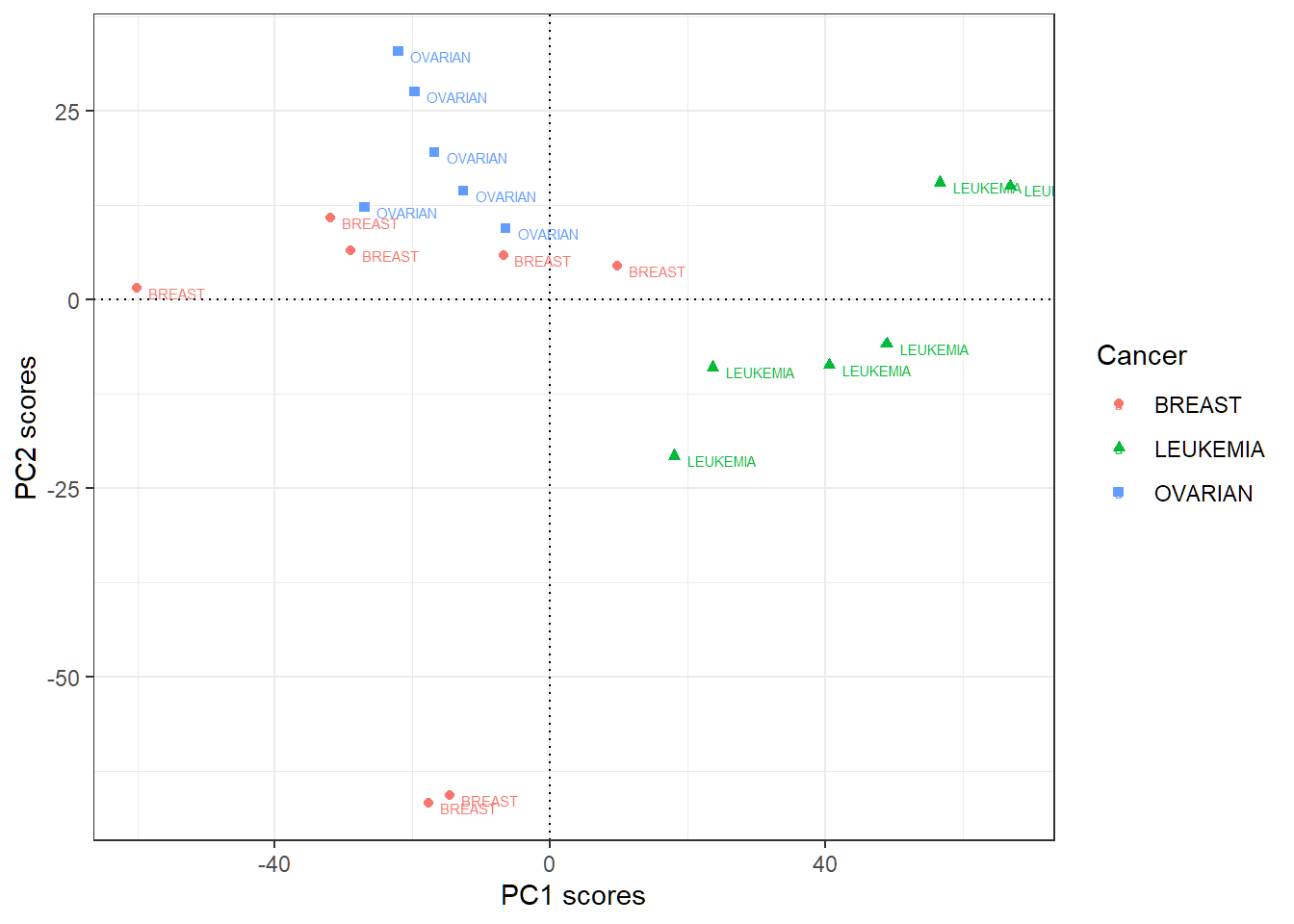
Sparse principal component analysis (SPCA): software implementation
R software and commands
R libraries and commands needed:
- R library
elasticnetand its commandspcafor sparse PCA
Command spca
spca{elasticnet} implements SPCA using an alternating minimization algorithm to minimize the SPCA objective function. Its basic syntax is:
spca(x, K, para, type=c("predictor","Gram"),
sparse=c("penalty","varnum"), use.corr=FALSE,
lambda=1e-6,
max.iter=200, trace=FALSE, eps.conv=1e-3)x: A matrix. It can be the predictor matrix or the sample covariance/correlation matrix.K: Number of principal components to compute.para: A vector of lengthKwith all positive elements.
Command spca
Basic syntax:
spca(x, K, para, type=c("predictor","Gram"),
sparse=c("penalty","varnum"), use.corr=FALSE,
lambda=1e-6,
max.iter=200, trace=FALSE, eps.conv=1e-3)type: Iftype="predictor",xis the predictor matrix. Iftype="Gram", user needs to provide the sample covariance or correlation matrix.sparse: Ifsparse="penalty",parais a vector of \(l_1\)- penalty tuning parameters. Ifsparse="varnum", each element ofparadefines the number of nonzero entries for a loading vector. This option is convenient in practice.
Command spca
Basic syntax:
spca(x, K, para, type=c("predictor","Gram"),
sparse=c("penalty","varnum"), use.corr=FALSE,
lambda=1e-6,
max.iter=200, trace=FALSE, eps.conv=1e-3)lambda: Quadratic penalty, i.e., \(l_2\)-penalty tuning parameter. Default value is1e-6.use.corr: Perform SPCA on the correlation matrix? This option is only effective when the argument type is set to be “data”.max.iterandeps.conv: Maximum number of iterations and convergence criterion for optimization, respectively.trace: IfTRUE, print out progress of optimization.
Command spca
spca returns an “spca” object that has the following components:
loadings: The matrix of loading vectors for theKsparse PCs (SPCs). The \(i\)-th column ofloadingsis for SPC \(i\).pev: Percentage of explained variance for each of theKsparse PCs. The \(i\)-th entry ofpevis for SPC \(i\).var.all: Total variance of the predictors.- The \(i\)-th column of
x%*%loadingsis the score vector for SPC \(i\) and pairs with the \(i\)-th column ofloadings
Sparse principal component analysis: Example 1
pitprops data
The
pitpropsdata is a correlation matrix that was calculated from 180 observations on 13 featuresOriginal data matrix is not available here
This is a classical example showing the difficulty of interpreting principal components
Corrrelation matrix
> library(corrplot); library(elasticnet); data(pitprops)
> corrplot(pitprops,method="shade",tl.cex=0.7,mar=rep(1,4))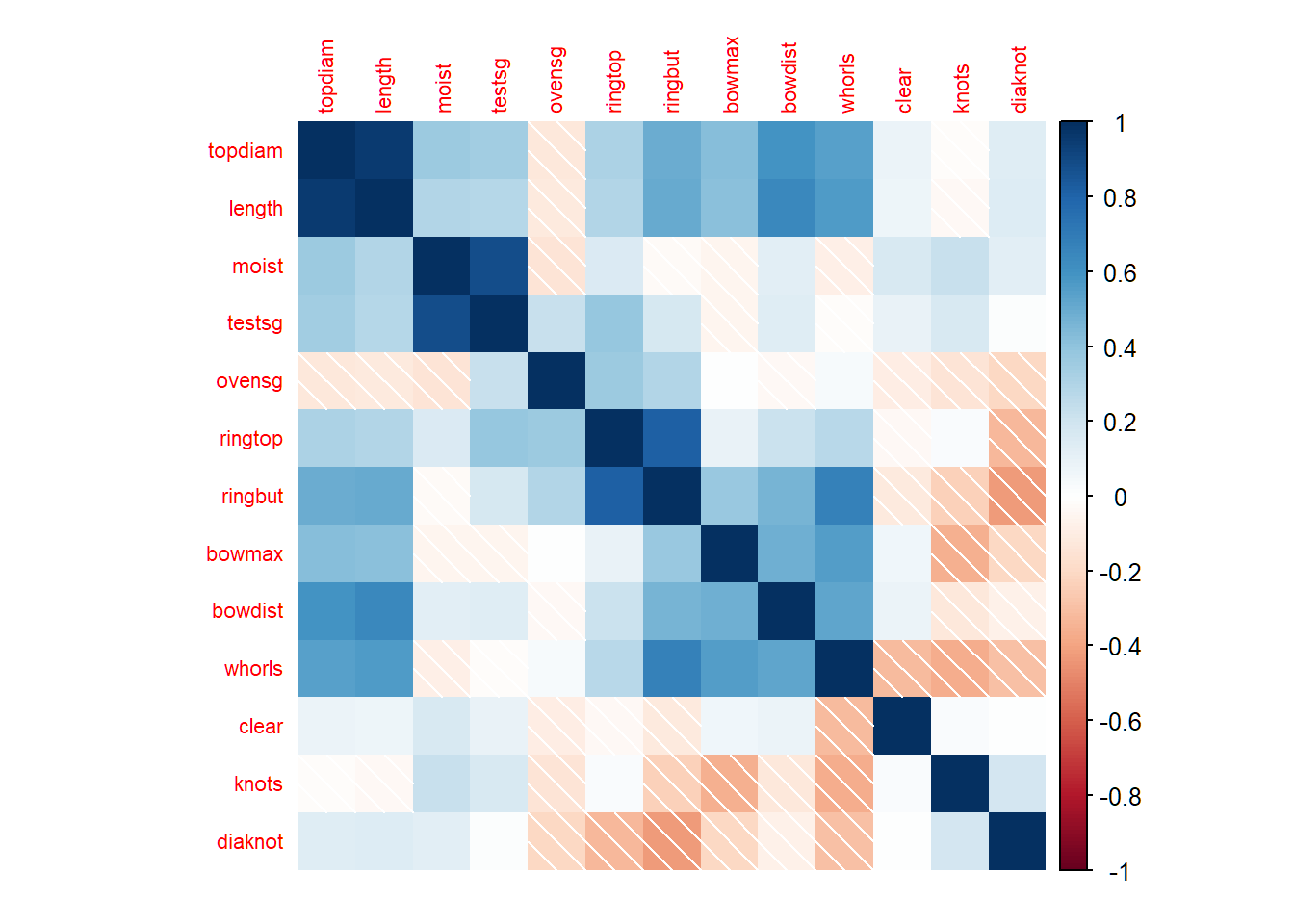
SPCA via tuning parameters
The first 6 sparse PCs are obtained by setting tuning parameters:
> library(elasticnet); data(pitprops)
> out1=elasticnet::spca(pitprops,K=6,type="Gram",
+ sparse="penalty",lambda=1e-6,
+ para=c(0.06,0.16,0.1,0.5,0.5,0.5))
> # loading vectors
> out1$loadings
PC1 PC2 PC3 PC4 PC5 PC6
topdiam -0.4773598 0.00000000 0.00000000 0 0 0
length -0.4758876 0.00000000 0.00000000 0 0 0
moist 0.0000000 0.78471386 0.00000000 0 0 0
testsg 0.0000000 0.61935898 0.00000000 0 0 0
ovensg 0.1765675 0.00000000 0.64065264 0 0 0
ringtop 0.0000000 0.00000000 0.58900859 0 0 0
ringbut -0.2504731 0.00000000 0.49233189 0 0 0
bowmax -0.3440474 -0.02099748 0.00000000 0 0 0
bowdist -0.4163614 0.00000000 0.00000000 0 0 0
whorls -0.4000254 0.00000000 0.00000000 0 0 0
clear 0.0000000 0.00000000 0.00000000 -1 0 0
knots 0.0000000 0.01333114 0.00000000 0 -1 0
diaknot 0.0000000 0.00000000 -0.01556891 0 0 1
> # adjusted percentage of explained variance
> out1$pev
[1] 0.28034916 0.13965535 0.13298210 0.07444957 0.06801883
[6] 0.06227288SPCA via sparsity levels
The first 6 sparse PCs are obtained by setting numbers of nonzero entries for sparse loading vectors:
> library(elasticnet); data(pitprops)
> out2<-elasticnet::spca(pitprops,K=6,type="Gram",
+ sparse="varnum",para=c(7,4,4,1,1,1))
> out2$loadings
PC1 PC2 PC3 PC4 PC5 PC6
topdiam -0.4774878 0.00273577 0.00000000 0 0 0
length -0.4691409 0.00000000 0.00000000 0 0 0
moist 0.0000000 0.78520583 0.00000000 0 0 0
testsg 0.0000000 0.61854739 0.00000000 0 0 0
ovensg 0.1797963 0.00000000 0.65551907 0 0 0
ringtop 0.0000000 0.00000000 0.58924631 0 0 0
ringbut -0.2898492 0.00000000 0.46990985 0 0 0
bowmax -0.3425338 -0.02904206 -0.04762634 0 0 0
bowdist -0.4138718 0.00000000 0.00000000 0 0 0
whorls -0.3833453 0.00000000 0.00000000 0 0 0
clear 0.0000000 0.00000000 0.00000000 -1 0 0
knots 0.0000000 0.00000000 0.00000000 0 -1 0
diaknot 0.0000000 0.00000000 0.00000000 0 0 1
> # adjusted percentage of explained variance
> out2$pev
[1] 0.28171026 0.13933060 0.13067145 0.07439423 0.06845471
[6] 0.06327273- PC1 loading vector has 7 nonzero entries
Sparse principal component analysis: Example 2
Human cancer microarray data
Human cancer data:
- 6830 genes (i.e., \(p=6830\) features) and their expressions
- 64 samples (i.e., \(64\) cancer cell lines)
- 14 different cancer types; information on cancer types at wiki-NCI60
We will pick all genes for 3 caner types “BREAST”, “OVARIAN” and “LEUKEMIA”, and this gives a data matrix \(\mathbf{X} \in \mathbb{R}^{19 \times 6830}\)
SPCA on standardized data
- SPCA will be implemented with tuning parameter \(\lambda=10^{-6}\) for \(l_2\)-penalty and tuning parameters \(\lambda_{1,j}=10^{-6}\) for \(1 \le j \le 18\) for \(l_1\)-penalty
> library(ISLR)
> nci.labs = NCI60$labs; nci.data = NCI60$data
> # pick 3 cancer types: BREAST, OVARIAN, LEUKEMIA
> labs1=nci.labs[nci.labs %in% c('BREAST','OVARIAN','LEUKEMIA')]
> data1=nci.data[nci.labs %in% c('BREAST','OVARIAN','LEUKEMIA'),]
> Xsd = scale(data1,center=TRUE,scale=TRUE)
> # para: amount of l1-penalty if sparse="penalty"
> # lambda: amount of l2-penalty
> # K: number of sparse PCs to be computed
> library(elasticnet)
> resB =elasticnet::spca(Xsd,K=18,para=rep(1e-6,18),
+ type=c("predictor"),sparse=c("penalty"),lambda=1e-6,
+ max.iter=200, eps.conv=1e-3)
> dim(resB$loadings)
[1] 6830 18First 4 loading vectors
Obtain histograms for first 4 sparse loading vectors:
> library(ggplot2); library(reshape2)
> # the first 4 sparse loading vectors
> FourLD = resB$loadings[,1:4]
> # change column names
> colnames(FourLD) = paste("SPC",1:4,sep="")
> colnames(FourLD)=paste(colnames(FourLD),"loadings",sep=" ")
> # melt and plot
> dstack <- melt(FourLD)
> oLV=ggplot(dstack,aes(x = value))+geom_histogram(bins = 25)+
+ facet_wrap(~Var2,scales="free")+theme_bw()First 4 sparse loading versions
Histograms for first 4 sparse loading vectors
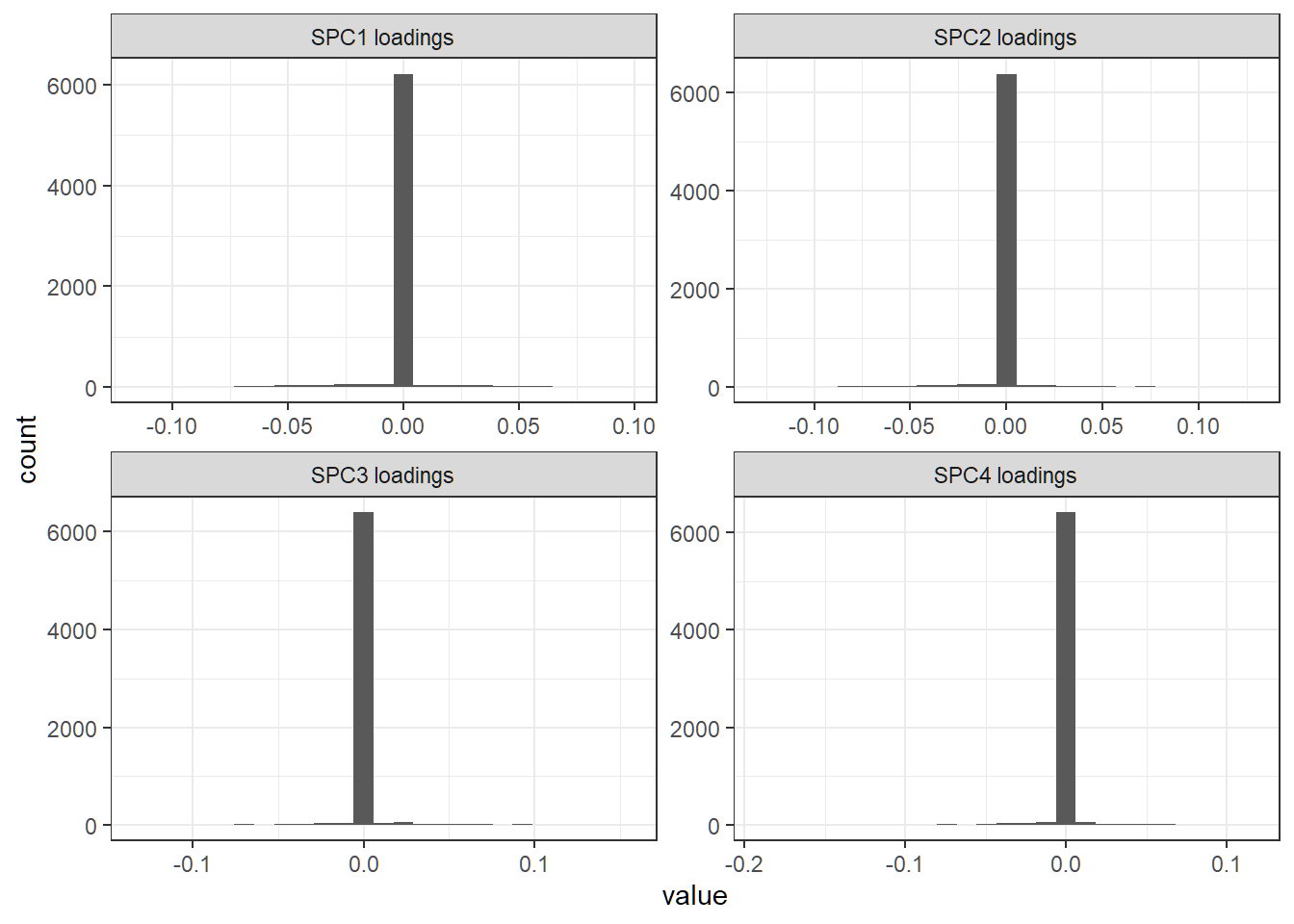
Dominant features
Dominant features (i.e., genes) in sparse direction for SPC1:
> # FourLD has the first 4 sparse loading vectors
> # as its 4 columns
> LV1 = FourLD[,1] # first sparse loading vector
> # all dominant features as those with nonzero loadings
> DF1 = which(abs(LV1)>0)
> length(DF1)
[1] 680
> # dominant features as those with large loadings
> DF2= which(abs(LV1)>
+ quantile(abs(LV1[abs(LV1)>0]),probs=c(0.95)) )
> length(DF2)
[1] 34- Features having nonzero or relatively large (in absolute value) loadings for a loading vector are dominant ones in that direction
- Dominant features can be tracked and identified by their indices or variable names
Adjusted PVE and CPVE
> pve=resB$pev # adjusted PVE; 18 sparse PCs
> # cumsum(pve) # adjusted CPVE
> par(mfrow=c(1,2),mar=c(9,4,1,3),mgp=c(1.8,0.6,0.3))
> plot(pve,type="o",ylab="Adjusted PVE",xlab="Sparse PC",col="blue")
> plot(cumsum(pve),type="o",ylab="Adjusted Cumulative PVE",
+ xlab="Number of first sparse PCs",col="blue")
> abline(h = cumsum(pve)[11], col="red", lty=2)
> abline(v = 11, col="red", lty=2)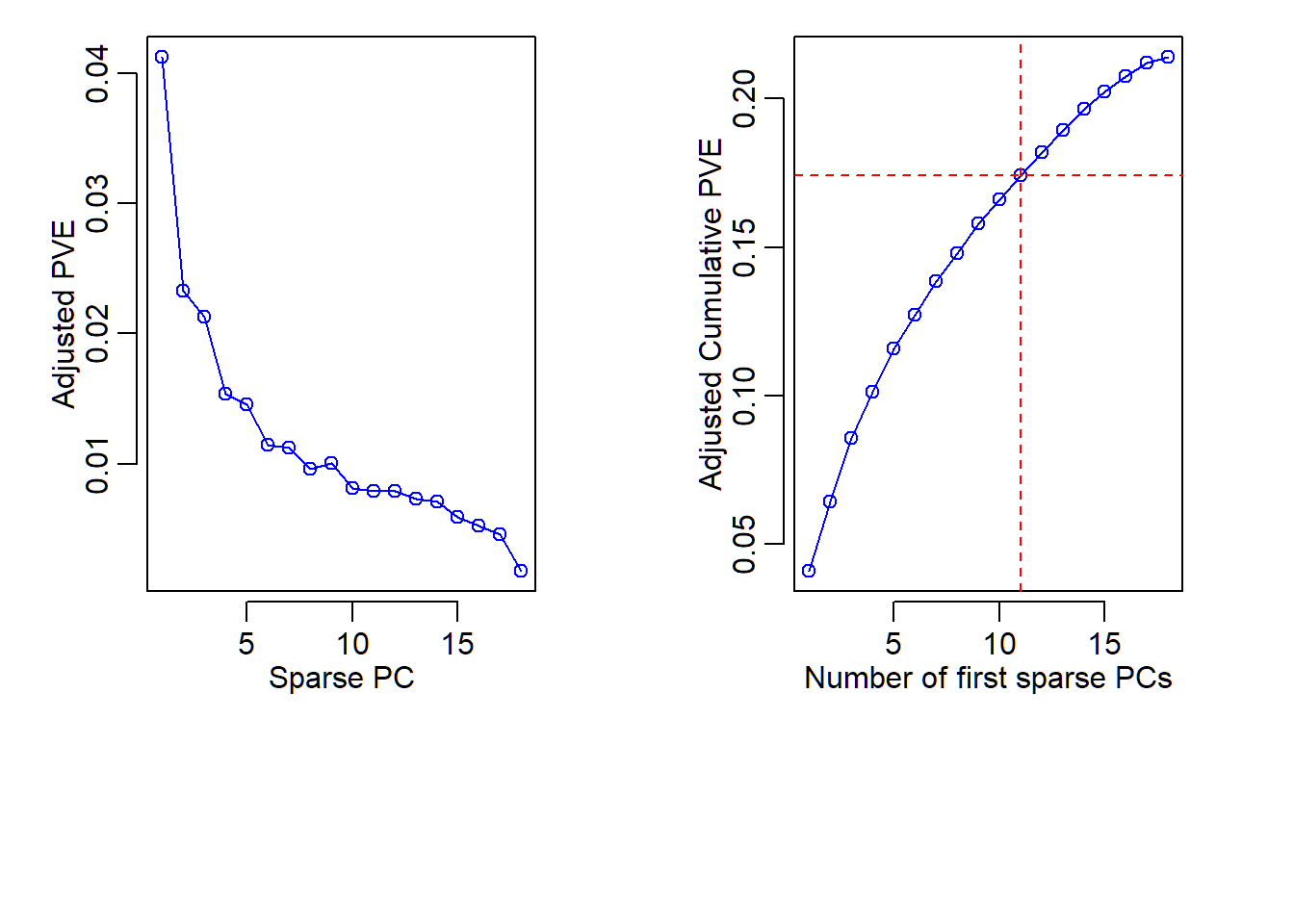
SPCA biplot
Codes to create a biplot for SPCA are bit long but very elementary:
> # get the first 2 loading vectors and score vectors
> sLVTwo=resB$loadings[,1:2]; sSVTwo= Xsd%*%sLVTwo
> # create color and shape scheme
> pch.group=rep(21,nrow(sSVTwo))
> pch.group[nci.labs1=='BREAST']=21;
> pch.group[nci.labs1=='OVARIAN']=22;
> pch.group[nci.labs1=='LEUKEMIA']=23
> col.group = rep("blue",nrow(sSVTwo))
> col.group[nci.labs1=='BREAST']="blue";
> col.group[nci.labs1=='OVARIAN']="green";
> col.group[nci.labs1=='LEUKEMIA']="cyan"
> # create plot
> par(mar=c(5,4.5,1,1),mgp=c(1.5,0.5,0))
> plot(sSVTwo[,1], sSVTwo[,2], xlab= "SPC1", ylab="SPC2",
+ col="black",pch=pch.group,bg=col.group,
+ las=1,asp=1,cex=0.8)
> # add center lines
> abline(v=0, lty=2, col="grey50")
> abline(h=0, lty=2, col="grey50")
> # add labels
> text(sSVTwo[,1],sSVTwo[,2],labels=nci.labs1,
+ pos=c(1,3,4,2),cex=.6)
> # ending positions of arrows
> l.x=100*sLVTwo[,1]; l.y=100*sLVTwo[,2]
> # draw arrows
> arrows(x0=0, x1=l.x, y0=0, y1=l.y,
+ col="red",length=0.1, lwd=1.5)SPCA biplot
- Hard to visualize sparse loadings due to too many features
- Scores show some patterns for different cancers (similar to those revealed by ordinary PCA scores)
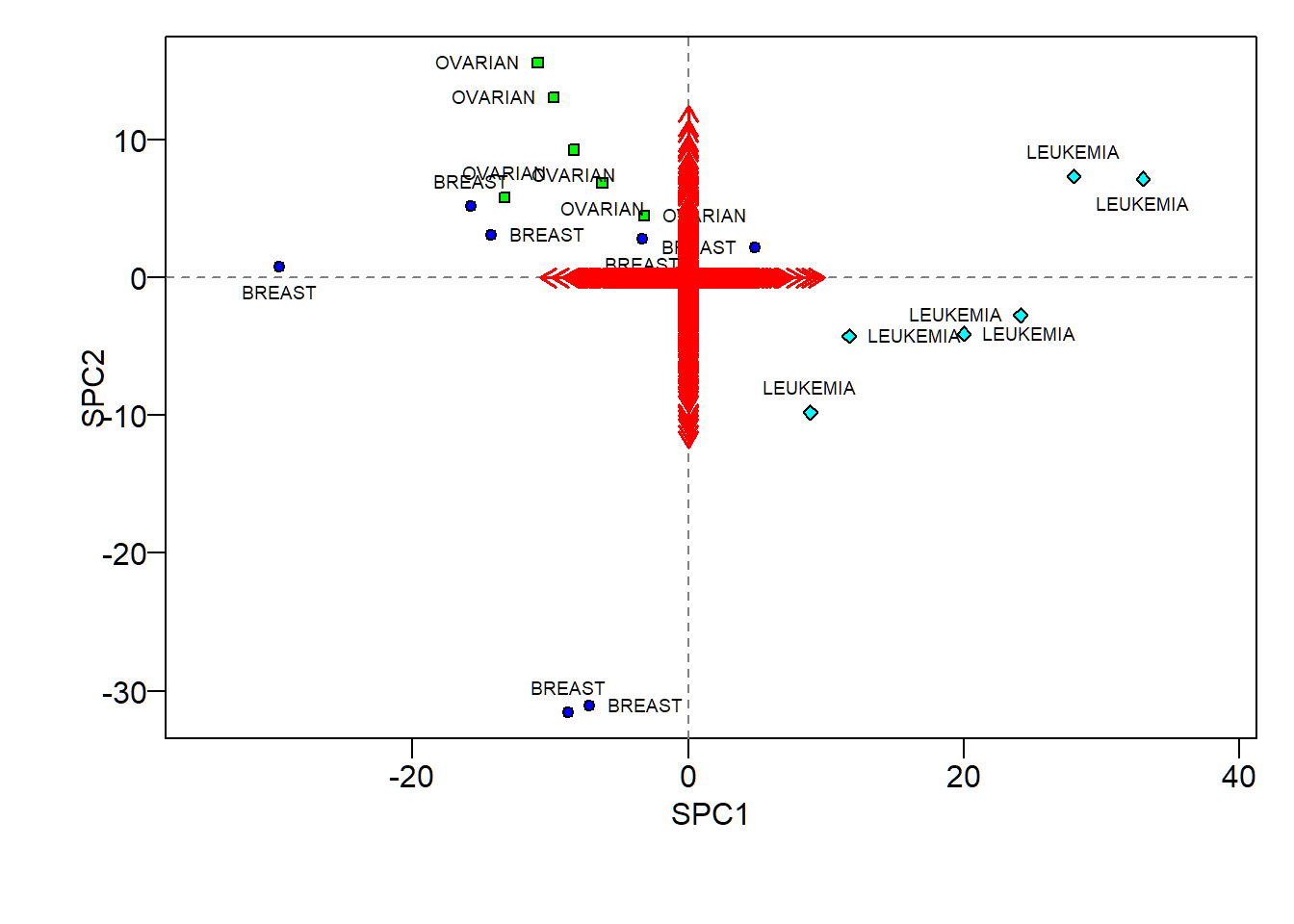
Caution
library(sparsepca) and library(elasticnet) both implement SPCA but library(sparsepca) may not adjusted PVE
> library(sparsepca)
> # `beta` for l2 penalty and `alpha` for l1 penalty
> resC =sparsepca::spca(Xsd, k=18, alpha=1e-6,beta=1e-6,
+ center=T,scale=T,max_iter=200,
+ tol=1e-05,verbose = FALSE)
> summary(resC)
PC1 PC2 PC3 PC4
Explained variance 1157.853 708.803 649.256 499.433
Standard deviations 34.027 26.623 25.481 22.348
Proportion of variance 0.170 0.104 0.095 0.073
Cumulative proportion 0.170 0.273 0.368 0.441
PC5 PC6 PC7 PC8 PC9
Explained variance 455.948 390.844 373.202 325.020 323.587
Standard deviations 21.353 19.770 19.318 18.028 17.989
Proportion of variance 0.067 0.057 0.055 0.048 0.047
Cumulative proportion 0.508 0.565 0.620 0.668 0.715
PC10 PC11 PC12 PC13 PC14
Explained variance 284.448 272.839 268.269 258.047 245.667
Standard deviations 16.866 16.518 16.379 16.064 15.674
Proportion of variance 0.042 0.040 0.039 0.038 0.036
Cumulative proportion 0.757 0.797 0.836 0.874 0.910
PC15 PC16 PC17 PC18
Explained variance 212.511 187.062 156.990 58.830
Standard deviations 14.578 13.677 12.530 7.670
Proportion of variance 0.031 0.027 0.023 0.009
Cumulative proportion 0.941 0.968 0.991 1.000
> # par(mfrow=c(1,2),mar=c(9,4,1,3),mgp=c(1.8,0.6,0.3))
> # plot(1:ncol(sResB),sResB[3,],type="o",ylab="PVE",xlab="Sparse PC",col="blue")
> # plot(sResB[4,],type="o",ylab="Cumulative PVE",xlab="Number of sparse PCs",col="blue")
> # abline(h = cumsum(pve)[11], col="red", lty=2)
> # abline(v = 11, col="red", lty=2)Appendix 0: Customized biplot
R software and commands
R libraries and commands needed:
- R library
ggbiplotand its commandggbiplotto customize biplot
Command ggbiplot
ggbiplot{ggbiplot} creates a biplot for PCA using ggplot2. Its basic syntax is:
ggbiplot(pcobj, choices = 1:2, scale = 1, pc.biplot =
TRUE, obs.scale = 1 - scale, var.scale = scale, groups =
NULL, labels = NULL, labels.size = 3, alpha = 1,
var.axes = TRUE, varname.size = 3,
varname.adjust = 1.5, varname.abbrev = FALSE)pcobj: an object returned by commandprcompchoices: which PCs to plot
Command ggbiplot
Basic syntax:
ggbiplot(pcobj, choices = 1:2, scale = 1, pc.biplot =
TRUE, obs.scale = 1 - scale, var.scale = scale, groups =
NULL, labels = NULL, labels.size = 3, alpha = 1,
var.axes = TRUE, varname.size = 3,
varname.adjust = 1.5, varname.abbrev = FALSE)scale: covariance biplot (scale = 1), form biplot (scale = 0). Whenscale = 1, the inner product between the variables approximates the covariance and the distance between the points approximates the Mahalanobis distance.obs.scale: scale factor to apply to observationsvar.scale: scale factor to apply to variables
Command ggbiplot
Basic syntax:
ggbiplot(pcobj, choices = 1:2, scale = 1, pc.biplot =
TRUE, obs.scale = 1 - scale, var.scale = scale, groups =
NULL, labels = NULL, labels.size = 3, alpha = 1,
var.axes = TRUE, varname.size = 3,
varname.adjust = 1.5, varname.abbrev = FALSE)pc.biplot: for compatibility with commandbiplot.princompgroups: optional factor variable indicating the groups that observations belong to. If provided, points will be colored according to groupslabels: optional vector of labels for the observationslabels.size: size of the text used for the labels
Command ggbiplot
Basic syntax:
ggbiplot(pcobj, choices = 1:2, scale = 1, pc.biplot =
TRUE, obs.scale = 1 - scale, var.scale = scale, groups =
NULL, labels = NULL, labels.size = 3, alpha = 1,
var.axes = TRUE, varname.size = 3,
varname.adjust = 1.5, varname.abbrev = FALSE)alpha: transparency value for the points (0= transparent,1= opaque)var.axes: draw arrows for the variables?varname.sizeandvarname.abbrev: size of the text for and whether or not to abbreviate variable namesvarname.adjust: adjustment factor the placement of the variable names; a value>= 1means farther from the arrow
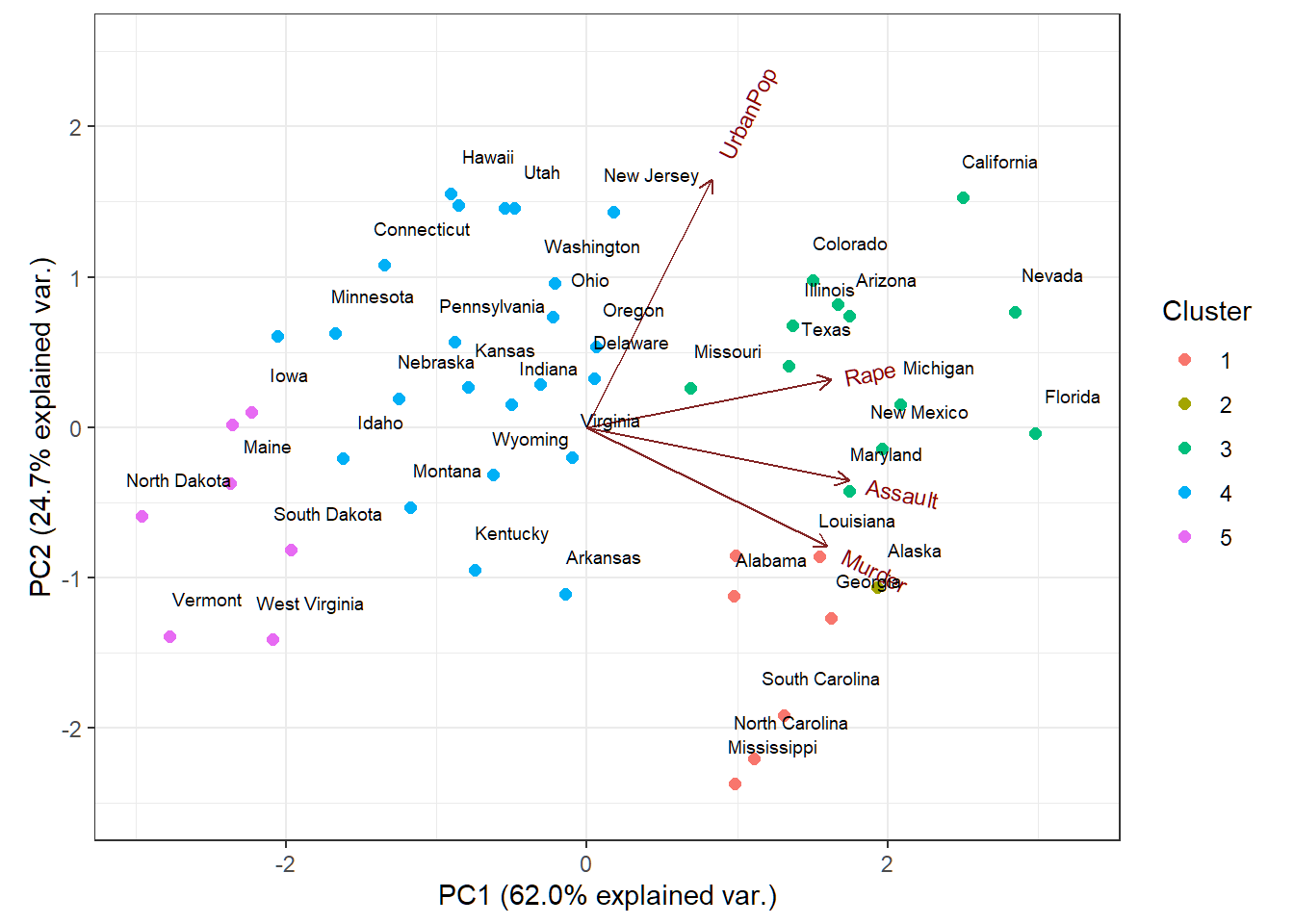
Biplot with clustering
Cluster memberships of US states superimposed on biplot of PCA of USArrests data
> library(ISLR)
> Xsd = scale(USArrests,center=TRUE,scale=TRUE)
> # hierarchical clustering
> hc=hclust(dist(Xsd), "ave"); hc5=cutree(hc, k=5)
> pcaXsd = prcomp(Xsd)
> # change orientation or loading and score vectors
> pcaXsd$rotation= -pcaXsd$rotation; pcaXsd$x= -pcaXsd$x
> # better biplot
> library(ggbiplot)
> cbiF= ggbiplot(pcaXsd,choices=1:2,obs.scale=1,var.scale=1)+
+ geom_point(aes(color=factor(hc5)),size=2)+
+ labs(colour="Cluster",shape="Cluster")+
+ theme(legend.direction = 'horizontal',
+ legend.position = 'top')+ylim(-2.5,2.5)+
+ theme_bw()+geom_text(label=rownames(USArrests),
+ nudge_x=0.25, nudge_y=0.25, cex=2.5, check_overlap=T)Biplot with clustering
Cluster memberships superimposed on biplot
Appendix 1: command princomp{stats}
Command princomp{stats}
princomp{stats} performs a PCA on a given numeric data matrix and returns the results as an object of class princomp. Its basic syntax is:
princomp(x, cor = FALSE, scores = TRUE, covmat = NULL,
subset = rep_len(TRUE, nrow(as.matrix(x))),
fix_sign = TRUE, ...)x: a numeric matrix or data frame which provides the data for the principal components analysis.cor: a logical value indicating whether the calculation should use the correlation matrix or the covariance matrix. (The correlation matrix can only be used if there are no constant variables.)
Command princomp{stats}
Basic syntax:
princomp(x, cor = FALSE, scores = TRUE, covmat = NULL,
subset = rep_len(TRUE, nrow(as.matrix(x))),
fix_sign = TRUE, ...)scores: a logical value indicating whether the score on each principal component should be calculated.covmat: a covariance matrix, or a covariance list as returned bycov.wt(andcov.mveorcov.mcdfrom packageMASS). If supplied, this is used rather than the covariance matrix ofx.
Command princomp{stats}
Basic syntax:
princomp(x, cor = FALSE, scores = TRUE, covmat = NULL,
subset = rep_len(TRUE, nrow(as.matrix(x))),
fix_sign = TRUE, ...)fix_sign: Should the signs of the loadings and scores be chosen so that the first element of each loading is non-negative? The signs of the columns of the loadings and scores are arbitrary, and so may differ between different programs for PCA, and even between different builds of R:fix_sign=TRUEalleviates that....: arguments passed to or from other methods. Ifxis a formula, one might specifycororscores.
Command princomp{stats}
princompis a generic function with “formula” and “default” methods.- The calculation is done using the command
eigenon the correlation or covariance matrix, as determined bycor. A preferred method of calculation is to usesvdonx, as is done inprcomp. - Note that the default calculation uses divisor
n(the sample size) for the covariance matrix. - The
printmethod for these objects prints the results in a nice format, and theplotmethod produces a scree plot (screeplot). There is also abiplotmethod.
Command princomp{stats}
princomp returns a list with class “princomp” containing the following components:
sdev: the standard deviations of the principal components.loadings: the matrix of variable loadings (i.e., a matrix whose columns contain the eigenvectors). This is of class “loadings”; seeloadingsfor itsprintmethod.center: the means that were subtracted.scale: the scalings applied to each variable.n.obs: the number of observations.
Command princomp{stats}
scores: ifscores=TRUE, the scores of the supplied data on the principal components. These are non-null only ifxwas supplied, and ifcovmatwas also supplied it was a covariance list. For the formula method,napredict()is applied to handle the treatment of values omitted by thena.action.call: the matched call.na.action: If relevant.
License and session Information
> sessionInfo()
R version 3.5.0 (2018-04-23)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19041)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.1252
[2] LC_CTYPE=English_United States.1252
[3] LC_MONETARY=English_United States.1252
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods
[7] base
other attached packages:
[1] knitr_1.21
loaded via a namespace (and not attached):
[1] compiler_3.5.0 magrittr_1.5 tools_3.5.0
[4] htmltools_0.3.6 revealjs_0.9 yaml_2.2.0
[7] Rcpp_1.0.0 stringi_1.2.4 rmarkdown_1.11
[10] stringr_1.3.1 xfun_0.4 digest_0.6.18
[13] evaluate_0.12